Introduction
Palmer amaranth (Amaranthus palmeri S. Watson) is an annual weed species native to the area encompassing northwestern Mexico and the southern United States (Sauer Reference Sauer1957). Over the last two decades, A. palmeri has become one of the most widespread, troublesome, and economically damaging weed species in several annual crop systems, particularly soybean [Glycine max (L.) Merr.], maize (Zea mays L.), and cotton (Gossypium hirsutum L.), in several countries, including the United States, Argentina, Brazil, Uruguay, and Mexico (Culpepper et al. Reference Culpepper, Grey, Vencill, Kichler, Webster, Brown, York, Davis and Hanna2006, Reference Culpepper, Whitaker, MacRae and York2008; Gaines et al. Reference Gaines, Gancho Slavov, Hughes, Kuepper, Sparks, Oliva, Vila-Aiub, Garcia, Merotto and Neve2021; Heap Reference Heap2022; Küpper et al. Reference Küpper, Borgato, Patterson, Netto, Nicolai, de Carvalho and Christoffoleti2017; Morichetti et al. Reference Morichetti, Cantero, Núñez, Barboza, Amuchastegui and Ferrell2013; Ward et al. Reference Ward, Webster and Steckel2013). Depending on the infestation level, yield losses from A. palmeri are reported to be from ∼10% to 100% (Bensch et al. Reference Bensch, Horak and Peterson2003; Van De Stroet Reference Van De Stroet2018). In addition, the presence of A. palmeri has been reported in more than 30 countries, globally (USDA-APHIS 2020). However, in many of these countries, it is present as a casual alien and may not be fully established (USDA-APHIS 2020).
Amaranthus palmeri is a new exotic plant species in the Republic of South Africa (RSA). It was first detected in the 2017 to 2018 summer growing season on a farm in the Douglas district of the Northern Cape Province (29.05°S, 23.77°E) in a variety of crops grown on the farm: alfalfa (Medicago sativa L.), cotton, and maize. Amaranthus palmeri was found growing together with another Amaranthus species, locally called Cape pigweed or common pigweed (Amaranthus hybridus L. ssp. hybridus var. hybridus) (Botha Reference Botha2010; Sukhorukov et al. Reference Sukhorukov, Kushumina, Reinhardt, Bezuidenhout and Vorster2021). In addition, spiny amaranth (Amaranthus spinosus L.), which originated from tropical America (Botha Reference Botha2010), has been well established in the country for many decades. Hybridization between A. palmeri (as pollen source) and A. hybridus was estimated to occur at a frequency of <0.01% when they coexist within 3 m (Gaines et al. Reference Gaines, Ward, Bekun, Preston, Leach and Westra2012). The same authors found that the highest levels of hybridization (up to 0.4%) were attained between A. palmeri and A. spinosus, which is genetically the species most closely related to A. palmeri. The South African A. palmeri population in the Douglas district initially drew the attention of farm management, because it was difficult to control with glyphosate (a nonselective herbicide) in a glyphosate-tolerant cotton crop. Two other populations have since been confirmed in the country, and another in Botswana, a country bordering the RSA to the northeast (Sukhorukov et al. Reference Sukhorukov, Kushumina, Reinhardt, Bezuidenhout and Vorster2021). The authors have provided geographic locations of these populations (Sukhorukov et al. Reference Sukhorukov, Kushumina, Reinhardt, Bezuidenhout and Vorster2021). These populations represent the first records of A. palmeri in southern Africa. Depending on the rate of spread, these known nonnative A. palmeri populations and the extent of A. palmeri’s negative impact, the status of this weed in southern Africa may change to “invasive” (Sukhorukov et al. Reference Sukhorukov, Kushumina, Reinhardt, Bezuidenhout and Vorster2021).
Numerous factors have contributed to A. palmeri becoming a dominant and difficult-to-control weed species, including its rapid growth rate (Horak and Loughin Reference Horak and Loughin2000), high fecundity (Bond et al. Reference Bond, Oliver and Stephenson2006; Burke et al. Reference Burke, Schroeder, Thomas and Wilcut2007; Horak and Loughin Reference Horak and Loughin2000; Keeley et al. Reference Keeley, Carter and Thullen1987; Sellers et al. Reference Sellers, Smeda, Johnson, Kendig and Ellersieck2003), genetic diversity (Küpper et al. Reference Küpper, Manmathan, Giacomini, Patterson, McCloskey and Gaines2018a), ability to tolerate adverse conditions, and ability to evolve resistance to multiple herbicides (Burke et al. Reference Burke, Schroeder, Thomas and Wilcut2007; Liphadzi and Dille Reference Liphadzi and Dille2006; Shyam et al. Reference Shyam, Borgato, Peterson, Dille and Jugulam2021). Currently, there are 70 confirmed cases of A. palmeri resistance to various herbicides reported in the United States, Argentina, Brazil, Israel, Mexico, and Spain (Heap Reference Heap2022). The first case of herbicide resistance in A. palmeri was recorded in 1989 for trifluralin (SOA Group 3) in cotton and soybean in the United States (Heap Reference Heap2022). The first case of resistance to glyphosate (Group 9) was reported in 2004 from cotton production in the state of Georgia, USA (Culpepper et al. Reference Culpepper, Grey, Vencill, Kichler, Webster, Brown, York, Davis and Hanna2006; Heap Reference Heap2022; Sosnoskie et al. Reference Sosnoskie, Kichler, Wallace and Culpepper2011). Amaranthus palmeri populations resistant to glyphosate were shown to require from 1.5 to 115 times the rate of glyphosate to achieve 50% control compared with susceptible plants (Norsworthy et al. Reference Norsworthy, Griffith, Scott, Smith and Oliver2008; Steckel et al. Reference Steckel, Main and Ellis2008).
The most problematic cases of resistance in A. palmeri illustrate its ability to evolve resistance to herbicides of multiple sites of action (SOAs). The worst cases of multiple resistance in A. palmeri include resistance to herbicides representing five different SOAs in a biotype in Arkansas (USA) and to herbicides of six different SOAs in Kansas (USA) (Heap Reference Heap2022; Shyam et al. Reference Shyam, Borgato, Peterson, Dille and Jugulam2021). Between these two biotypes, a total of eight different SOAs are involved. The six way–resistant A. palmeri biotype in Kansas (USA) was found in a long-term conservation-tillage system where it was inadequately controlled by field rates of herbicides representing six SOAs: auxin mimic (2,4-D) and acetolactate synthase (ALS), photosystem II (PSII), 5-enolpyruvylshikimate-3-phosphate synthase (EPSPS), protoporphyrinogen oxidase (PPO), and 4-hydroxyphenylpyruvate dioxygenase (HPPD) inhibitors (Shyam et al. Reference Shyam, Borgato, Peterson, Dille and Jugulam2021). One of the most prevalent forms of multiple resistance in an A. palmeri population is resistance to EPSPS-inhibiting (glyphosate; Group 9) and ALS-inhibiting (Group 2) herbicides (Heap Reference Heap2022; Kohrt and Sprague Reference Kohrt and Sprague2017). Other herbicides for which A. palmeri biotypes were reported to have developed resistance include PS II inhibitors (Group 5), HPPD inhibitors (Group 27), PPO inhibitors (Group 14), auxin mimics (Group 4), very-long-chain fatty-acid (VLCFA) synthesis inhibitors (Group 15), glutamine synthetase inhibitor (glufosinate ammonium; Group 10), and/or microtubule assembly inhibitors (Group 3) (Heap Reference Heap2022).
There are three basic mechanisms by which weed species develop resistance to a herbicide: (1) target-site alteration, (2) enhanced metabolism of the herbicide, and (3) reduced herbicide access to the SOA within the plant cell (exclusion) (Küpper et al. Reference Küpper, Manmathan, Giacomini, Patterson, McCloskey and Gaines2018a; Sammons and Gaines Reference Sammons and Gaines2014; Sammons et al. Reference Sammons, Heering, DiNicola, Glick and Elmore2007; Ward et al. Reference Ward, Webster and Steckel2013). Avoidance was proposed as an additional mechanism whereby weeds avoid the toxic effect produced by the herbicide by inactivation of its biochemical ability (Sammons and Gaines Reference Sammons and Gaines2014). More than 200 distinct weed biotypes worldwide have developed herbicide resistance via target-site alteration (Devine and Shukla Reference Devine and Shukla2000). Recently, A. palmeri and a few other species—kochia [Bassia scoparia (L.) A.J. Scott], A. spinosus, waterhemp [Amaranthus tuberculatus (Moq.) Sauer], and Italian ryegrass [Lolium perenne L. ssp. multiflorum (Lam.) Husnot]—have been found with duplications of the EPSPS gene, hence creating much higher levels of EPSPS protein and thereby developing resistance to glyphosate (Koo et al. Reference Koo, Jugulam, Putta, Cuvaca, Peterson, Currie, Friebe and Gill2018a, Reference Koo, Molin, Saski, Jiang, Putta, Jugulam, Friebe and Gill2018b; Sammons and Gaines Reference Sammons and Gaines2014). Gaines et al. (Reference Gaines, Zhang, Wang, Bukun, Chisholm and Shaner2010) reported 5- to 160-fold increases in EPSPS gene copy number and corresponding increases in EPSPS protein levels in a glyphosate-resistant A. palmeri population compared with a susceptible population. In addition, certain populations of glyphosate-resistant A. palmeri were shown to have target-site resistance via mutation(s) in the EPSPS gene (Dominguez-Valenzuela et al. Reference Dominguez-Valenzuela, Gherekhloo, Fernández-Moreno, Cruz-Hipolito, Alcántara-de la Cruz, Sánchez-Gonzáles and De Prado2017; Kaundun et al. Reference Kaundun, Jackson, Hutchings, Galloway, Marchegiani, Howell, Carlin, McIndoe, Tuesca and Moreno2019).
In this study, we present the first characterization of an A. palmeri population found in the Douglas district of the RSA in terms of its sensitivity to key herbicides in a controlled environment, as well as molecular assessment of the possible mechanisms of resistance to herbicides with selected SOAs. The information presented is intended to support development of the best strategies for A. palmeri management and to minimize further spread in the RSA and further afield in southern Africa.
Materials and Methods
Materials
Seeds from a suspected glyphosate-resistant A. palmeri population (referred to hereafter as “AMAPA-NC” for “Amaranthus palmeri in Northern Cape Province”) were collected in April 2018 from a cotton field in the Douglas district (29.05°S, 23.77°E), Northern Cape Province, RSA. For the testing of this population’s susceptibility to various herbicides in a controlled environment, two herbicide-susceptible A. palmeri lines were used: AMAPA-S1 from Shickley, NE, USA (collected in 2011) and AMAPA-S2 from Macon, GA, USA (collected in 2006). Published GenBank sequences of susceptible A. palmeri were used for target-gene sequence analysis.
Herbicide Active Ingredients
Table 1 lists each herbicide used in the current study along with its trade name, HRAC’s herbicide SOA group number (HRAC 2021), and recommended label rate. All trade names and label rates provided in Table 1 are based on their registration in Germany, where plant assays were conducted. The status of registration of these active ingredients (trade names may vary for a particular active ingredient) in the RSA is also indicated.
Table 1. Pre- and postemergence herbicides used in this study. a
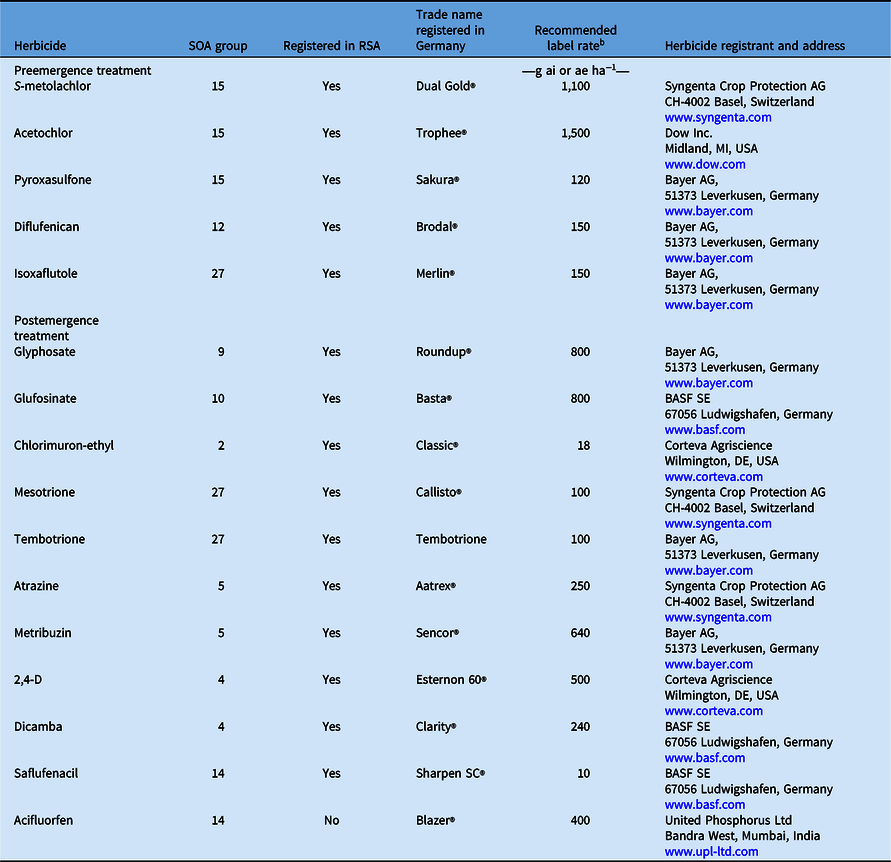
a Abbreviation: SOA, site of action; RSA, Republic of South Africa.
b Label rate registered in Germany.
Plant Growth Conditions
Approximately 10 to 20 A. palmeri seeds were directly sown into 8-cm-diameter Jiffy pots (Jiffy Products International B.V., Zwijndrecht, Netherlands) containing commercial potting soil at Bayer AG’s controlled environment facility in Frankfurt, Germany, during 2019. The emerged seedlings were thinned to maintain 5 plants per pot for postemergence herbicide treatments. The pots were maintained under greenhouse conditions of 28/22 C and 16/8 h day/night, respectively, with 50% relative humidity. The pots were irrigated, and plants were fertilized as necessary.
Evaluation of Pre- and Postemergence Herbicides
The experimental design involved 10 and 5 plants per pot for preemergence and postemergence herbicide treatments, respectively, each with three replicates per treatment. Individual preemergence herbicides were applied to the soil surface 3 d after A. palmeri seeds were sown in the pots, except for the untreated controls. These untreated controls also served to assess seed dormancy and viability. Individual postemergence herbicide treatments were applied when A. palmeri seedlings were at the 4-leaf stage. Individual herbicides were applied using a stationary research sprayer (Höchst AG, Höchst, Germany) calibrated to deliver a spray volume of 300 L ha−1 at a pressure of 200 kPa through a TeeJet® 8001EVS nozzle (TeeJet Technologies GmbH, Schorndorf, Germany). Herbicide product information and recommended label rates (1×) are presented in Table 1. Dose–response assays were conducted for each herbicide at seven different rates: 1/16×, 1/8×, 1/4×, 1/2×, 1×, 2×, and 4×, where 1× is the label rate. In addition, untreated controls were included for comparison. Efficacy and visual injury ratings were assessed at 25 d after herbicide treatment (DAT) for preemergence herbicides, with 0% indicating no damage and 100% indicating full damage, and at 16 DAT for postemergence herbicides, with 0% indicating no survival and 100% indicating complete survival.
Statistical Analysis
Dose–response analysis was conducted using the drc package in R (Knezevic et al. Reference Knezevic, Streibig and Ritz2007; R Core Team 2015) following Seefeldt et al. (Reference Seefeldt, Jensen and Fuerst1995). Survival data (proportion) were analyzed using the three-parameter log-logistic model in the drc package:
where y is survival, x is the herbicide dose, D is the upper limit, b is the slope, and LD50 is the dose causing 50% reduction in survival.
Assessment of Target-Site Mutations
Assessment of EPSPS, ALS, and PPO Target-Site Mutations
A leaf segment of approximately 0.5 cm2 was sampled from each of 20 A. palmeri and 10 A. hybridus individual plants directly from the field in the Douglas district, Northern Cape Province, RSA, and placed in a Costar 96-well block (Thermo Fisher Scientific, Johannesburg, RSA). These samples were frozen in liquid nitrogen until analysis at the University of Pretoria for potential mutations in EPSPS, ALS, and PPX2 (protoporphyrinogen oxidase 2 that is involved in conferring resistance to PPO inhibitors). For DNA extraction, leaf tissues were frozen in liquid nitrogen and ground into a fine powder using a sterile mortar and pestle. Genomic DNA was extracted using the Quick DNA™ Plant/Seed Miniprep Kit (Zymo Research, Inqaba, RSA) according to the manufacturer’s protocol. DNA quantification was done using a Nanodrop™ 2000 spectrophotometer (Thermo Fisher Scientific) and quality checked using 1% agarose gel electrophoresis. Target sites were amplified by polymerase chain reaction (PCR) in a Boeco TC-PRO (Boeco, Hamburg, Germany) thermocycler. Each PCR reaction contained 1× DreamTaq PCR Master Mix (Thermo Fisher Scientific), 400 nM each of the forward and reverse primers (Whitehead Scientific, Johannesburg, RSA), 50 ng of gDNA, and 10 µl of dH2O to a total volume of 25 µl. Thermoprofile conditions consisted of initial denaturation at 94 C for 5 min for all three genes tested; followed by 30 cycles of denaturation at 94 C for 30 s for all three genes (see Table 2), annealing (60 C for 30 s for EPSPS, 59 C for 30 s for ALS, and 62 C for 30 s to detect ΔG210 in PPX2), and extension at 72 C for 1 min for all three genes; and a final extension at 72 C for 10 min. The PCR products were confirmed by 1% agarose gel electrophoresis, and the desired DNA bands were excised and purified using a Zymoclean™ Gel DNA Recovery kit (Zymo Research, Inqaba, RSA). Purified PCR products of EPSPS and ALS were sequenced directly using Sanger sequencing at the African Centre for Gene Technologies (ACGT) sequencing facility at the University of Pretoria, Hatfield, RSA. Analysis and alignment of the sequences was carried out using CLC Genomic Workbench 8.0.1 (CLC Bio, Aarhus, Denmark). For the PPX2 amplified fragment, purified PCR products were ligated into the linearized pMiniT 2.0 vector using an NEB PCR cloning kit (New England Biolabs, Inqaba, RSA) and grown on the stable outgrowth medium provided with the kit at 37 C for 60 min with shaking at 250 rpm. The outgrowth was spread onto Luria Broth (LB) 100 µg ml−1 ampicillin plates (1% tryptone, 10 g L−1 yeast extract, 0.5% NaCl, and 1.5% agar; Sigma-Aldrich, St Louis, MO, USA) and incubated at 37 C overnight. The insert DNA was screened by colony PCR. Plasmid DNA from PCR-confirmed colonies was extracted using a Qiagen™ Miniplasmid purification kit (Qiagen, Hilden, Germany) and sequenced using Sanger sequencing at the ACGT DNA sequencing facility at the University of Pretoria.
Table 2. Target genes, their targeted sites of action, and polymerase chain reaction (PCR) primers used in the analysis.

For the Arg-98-Met and Arg-98-Gly mutation screening in the PPX2 target site, a dCAPS (derived cleaved amplified polymorphic sequences) assay developed by Giacomini et al. (Reference Giacomini, Umphres, Nie, Mueller, Steckel, Young, Scott and Tranel2017) was used. The dCAPS assay allows for rapid genotyping, by gel electrophoresis, of the presence or absence of two Arg-98 mutations based on restriction enzyme digestion of PCR fragments. The dCAPS involved a nested PCR carried out with an initial amplification using primers dCAPS-98-F and G210-R to amplify a 1,600-bp product (Table 2). A second PCR was carried using the dCAPS primer dCAPS-98-F and the reverse primers dCAPS-98-R1 and dCAPS-98-R2, which are specific to Arg-98-Met and Arg-98-Gly, respectively (Table 2), to amplify ∼500-bp fragments. The PCR reactions consisted of 1× DreamTaq master mix (Thermo Fisher Scientific), 400 nM of each primer (Integrated DNA Technology, Capetown, RSA), 9.5 µl of dH2O, and 20 to 50 ng of gDNA to a total volume of 20 µl. Thermoprofile conditions were as follows: initial denaturation at 94 C for 5 min; 30 cycles of denaturation at 94 C for 30 s, annealing at 56 C for 30 s, and elongation at 72 C for 1 min followed by final elongation at 72 C for 10 min; and hold at 4 C for 59 min using a Boeco TC-PRO thermocycler. The resulting PCR product was mixed with 1 unit of KpnI restriction enzyme to specifically detect the Arg-98-Met mutation or with HindIII to specifically detect the Arg-98-Gly mutation in a 1× FastDigest buffer (Thermo Fischer Scientific). Negative controls containing the PCR products and 1× FastDigest Buffer without the restriction enzyme were also prepared for all samples. All reactions were incubated at 37 C for 2 h for complete digestion. The digested reactions were analyzed by agarose (4%) gel electrophoresis using the following criteria: fully digested products were scored as wild type, partially digested were scored as heterozygous, and undigested products were scored as homozygous for the specific targeted mutation.
Assessment of EPSPS Copy Number
The same DNA samples used for target-gene sequencing described were tested for the relative copy number of EPSPS using quantitative real-time PCR (CFX96 Touch™, Bio-Rad, Johannesburg, RSA) as described by Gaines et al. (Reference Gaines, Zhang, Wang, Bukun, Chisholm and Shaner2010), with ALS used as a single-copy reference. Relative quantification was carried out as described by Gaines et al. (Reference Gaines, Zhang, Wang, Bukun, Chisholm and Shaner2010) using a modification of the 2−ΔΔCt method (Ct = cycle threshold). Relative quantification was expressed as ΔCt = (Ct of EPSPS − Ct of ALS), and 2ΔCt was calculated to obtain a relative increase in EPSPS copy number.
Results and Discussion
Response of Amaranthus palmeri AMAPA-NC Population to Preemergence Herbicides
Progeny of AMAPA-NC survivors from a glyphosate-treated cotton field were tested for efficacy against five preemergence herbicides representing three diverse SOAs (represented by HRAC Groups 15, 12, and 27; Table 1) that were known to offer good residual control of Amaranthus species in the RSA.
Response to Group 15 Herbicides
Among the three VLCFA inhibitors (Group 15) tested, S-metolachlor and acetochlor belong to the chloroacetamide chemical family, while pyroxasulfone belongs to the pyrazole class (HRAC 2021). Greenhouse screening of these herbicides on the A. palmeri population revealed differential efficacies, with S-metolachlor showing markedly reduced control (60% control) at the 1× (label) rate of 1,100 g ae ha−1; Table 3; Figure 1A) at 16 DAT, while acetochlor and pyroxasulfone were effective with 100% and 95% control, respectively, at the 1× rate (1,500 and 120 g ae ha−1, respectively) (Table 3). Interestingly, all three herbicides were effective at both the 2× and 4× rates (Table 3). At the sublethal rates of 1/4× and 1/2×, similar trends were observed, with S-metolachlor providing 0% and 20% control, respectively, while acetochlor provided 97% control at both sublethal rates and pyroxasulfone gave 87% and 88% control, respectively (Table 3; Figure 1B). Based on these results, acetochlor and pyroxasulfone are still effective weed control options, while careful consideration should be given to diversify their use to minimize further selection for S-metolachlor resistance in this population.
Table 3. Preemergence herbicides assessed, site-of-action (SOA) group, and efficacy against an Amaranthus palmeri population (AMAPA-NC) in South Africa.

a 1× represents the label rate for each herbicide as registered in Germany. Values in parentheses indicate SE for each treatment.
b LD50 is the dose causing 50% reduction in survival. N/A, not applicable.
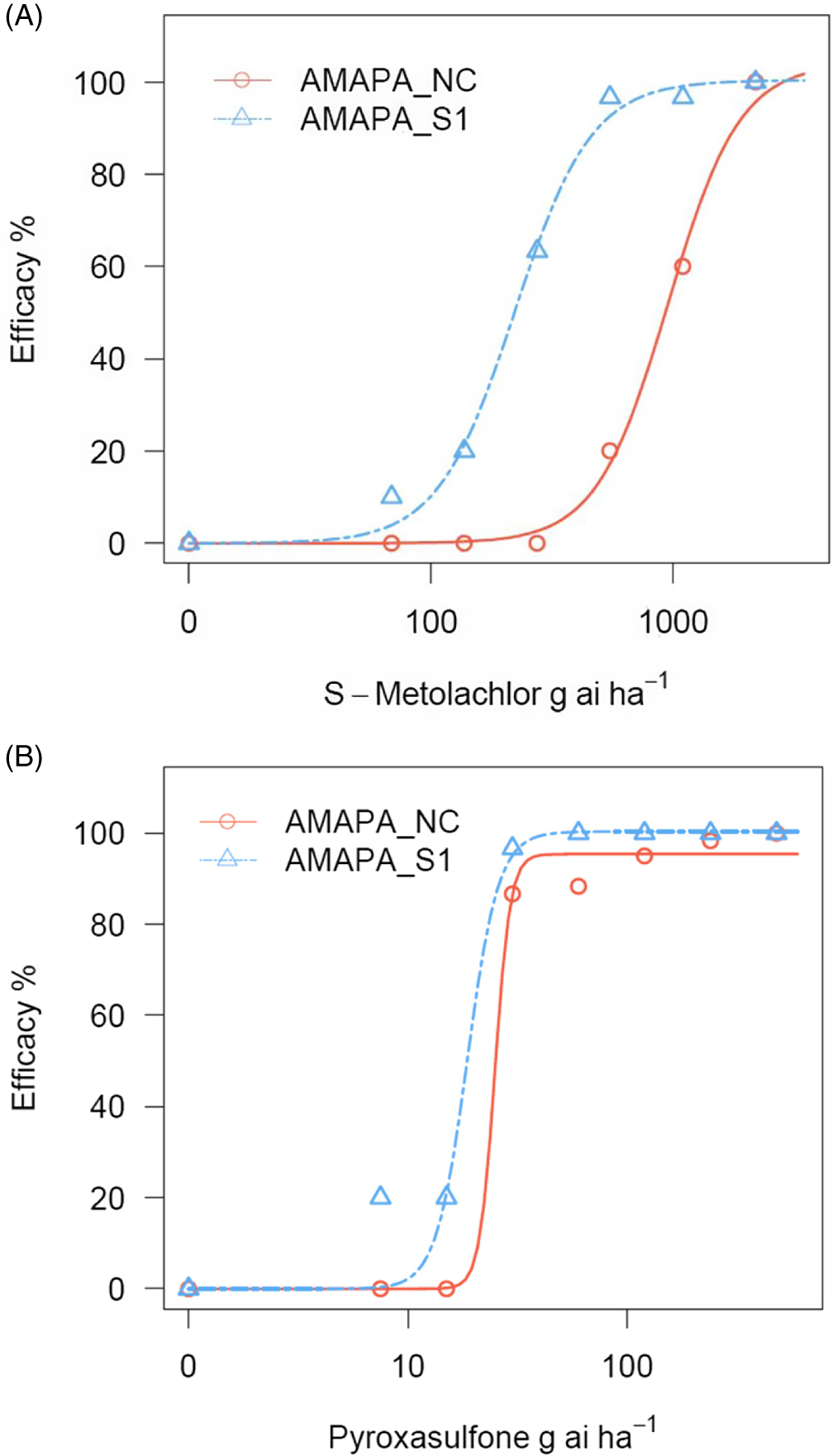
Figure 1. Percent efficacy at various dose levels of S-metolachlor (A) and pyroxasulfone (B) treatments (preemergence application) applied to the Amaranthus palmeri population AMAPA-NC found in the Republic of South Africa, and a sensitive population, AMAPA-S1 (Shickley, NE, USA). Efficacy was assessed at 25 d after treatment. Dose–response analysis was conducted using a three-parameter log-logistic model using the drc package in R software per Seefeldt et al. (Reference Seefeldt, Jensen and Fuerst1995).
The reduction in sensitivity to S-metolachlor in A. palmeri species (Brabham et al. Reference Brabham, Norsworthy, Houston, Varanasi and Barber2019; Rangani et al. Reference Rangani, Noguera, Salas-Perez, Benedetti and Roma-Burgos2021) and differential response to herbicides within SOA Group 15 were previously reported (Brabham et al. Reference Brabham, Norsworthy, Houston, Varanasi and Barber2019). Based on published studies, the mechanism of resistance to Group 15 herbicides in weedy species is largely attributed to non–target site resistance (NTSR), also referred to as metabolic resistance (Brabham et al. Reference Brabham, Norsworthy, Houston, Varanasi and Barber2019; Busi et al. Reference Busi, Porri, Gaines and Powles2018; Dücker et al. Reference Dücker, Zöllner, Lümmen, Ries, Collavo and Beffa2019a, 2019b; Rangani et al. Reference Rangani, Noguera, Salas-Perez, Benedetti and Roma-Burgos2021; Strom et al. Reference Strom, Hager, Seiter, Davis and Riechers2020, Reference Strom, Hager, Concepcion, Seiter, Davis, Morris, Kaundun and Riechers2021). It is postulated that target-site modification is an unlikely mechanism for resistance to Group 15 herbicides in crops and weedy species due to multiple SOAs represented by different enzymes involved in VLCFA synthesis (Busi et al. Reference Busi, Gaines, Vila-Aiub and Powles2014; Rangani et al. Reference Rangani, Noguera, Salas-Perez, Benedetti and Roma-Burgos2021). The NTSR mechanism imparting resistance to S-metolachlor in A. palmeri was previously reported to be due to glutathione-S-transferase (GST) detoxification in the roots (Rangani et al. Reference Rangani, Noguera, Salas-Perez, Benedetti and Roma-Burgos2021) as well as cytochrome P450–mediated metabolism (Rigon et al. Reference Rigon, Gaines, Küpper and Dayan2020; Strom et al. Reference Strom, Hager, Seiter, Davis and Riechers2020, Reference Strom, Hager, Concepcion, Seiter, Davis, Morris, Kaundun and Riechers2021). The differential specific activity of expressed Arabidopsis GST (AtGSTU19) gene homologues, which conjugate with acetochlor eight times higher compared with S-metolachlor, might explain the differences in sensitivity between these herbicides, but this mechanism needs further investigation (DeRidder et al. Reference DeRidder, Dixon, Beussman, Edwards and Goldsbrough2002). In addition, a study comparing tolerant and susceptible maize lines showed that the GST genes ZmGST5 and ZmGST6 were responsible for differential tolerance to S-metolachlor (Li et al. Reference Li, Gao, Xu, Pang, Liu, Wang and Tan2017). Similarly, S-metolachlor–resistant A. tuberculatus populations were shown to exhibit >2-fold GST activity compared with a susceptible population but much less than in tolerant maize. Conversely, cytochrome P450 activity in resistant A. tuberculatus was found to be >20-fold higher than in susceptible A. tuberculatus or maize (Strom et al. Reference Strom, Hager, Concepcion, Seiter, Davis, Morris, Kaundun and Riechers2021). Reduction in efficacy to S-metolachlor warrants careful resistance management of the A. palmeri AMAPA-NC population, because Group 15 herbicides are labeled for use in numerous crops for residual control of annual grasses and small-seeded broadleaf weeds.
Response to Isoxaflutole (Group 27) and Diflufenican (Group 12) Herbicides
Greenhouse screening of the A. palmeri AMAPA-NC population for efficacy against two other important preemergence herbicides, isoxaflutole and diflufenican, showed excellent control (∼100%) at various rates, including at sublethal rates as low as 1/4× the label rate (Table 3). Interestingly, at a very low sublethal rate of 1/8×, good control (90%) was achieved with both these herbicides (Table 3). Results indicated high sensitivity of this A. palmeri population under greenhouse conditions to these herbicides, which therefore provide an excellent preemergence herbicide treatment option for growers to manage this population. Herbicidal selectivity of diflufenican in weed species was shown to be primarily due to differential uptake and indirectly to translocation (Haynes and Kirkwood Reference Haynes and Kirkwood1992). However, it is known that isoxaflutole herbicide provides better control under field conditions when combined with an additional SOA that is efficacious against A. palmeri than as a stand-alone product (O’Brien et al. Reference O’Brien, Davis and Riechers2018; Spaunhorst and Johnson Reference Spaunhorst and Johnson2017).
Globally, there are two reported cases of resistance to isoxaflutole, namely, A. tuberculatus in the United States, which is also resistant to ALS inhibitors, atrazine, glyphosate, and mesotrione, and wild radish (Raphanus raphanistrum L.) in Australia, which is also resistant to diflufenican, chlorsulfuron, atrazine, fluridone, 2,4-D, mesotrione, and tembotrione (Heap Reference Heap2022). Hitherto, there has been no reported resistance to isoxaflutole in A. palmeri (Heap Reference Heap2022). Resistance to diflufenican herbicide was reported in four weed species—R. raphanistrum, Oriental mustard (Sisymbrium orientale L.), and capeweed [Arctotheca calendula (L.) Levyns] in Australia and eastern groundsel (Senecio vernalis Waldst. & Kit.) in Israel. Resistance to diflufenican in R. raphanistrum was reported to be due to its metabolism involving cytochrome P450s (Lu et al. Reference Lu, Yu, Han, Owen and Powles2020b). Three of the four diflufenican-resistant weed species (all except S. orientale) were also resistant to at least one additional SOA (Heap Reference Heap2022).
Response of Amaranthus palmeri AMAPA-NC Population to Postemergence Herbicides
Survival of A. palmeri plants after treatment with different postemergence herbicides was highly variable, indicating the considerable genetic variability of the AMAPA-NC population. Overall, 5 of the 11 postemergence herbicides tested showed less than 90% control at the label rate (Table 4). The lowest level of control was found for saflufenacil (33%) followed by glyphosate (60%) at their respective label rates; however, glufosinate, metribuzin, tembotrione, 2,4-D, and acifluorfen showed 100% control at the label rate. Note that acifluorfen, to our knowledge, is not yet registered in the RSA.
Table 4. Postemergence herbicides assessed, site-of-action (SOA) group, and efficacy against an Amaranthus palmeri population (AMAPA-NC) in South Africa.

a 1× represents the label rate for each herbicide as registered in Germany. Values in parentheses indicate SE for each treatment.
b LD50 is the dose causing 50% reduction in survival.
In response to glyphosate (EPSPS inhibitor; Group 9) treatment, A. palmeri AMAPA-NC population showed significantly reduced sensitivity at all doses tested, ranging from 80% control at 4× the label rate (800 g ae ha−1) to 60% control at the label rate. No control was observed at 2× the label rate and any of the sublethal doses tested (Table 4; Figure 2A). The unexpected trend between 1× (60% control) and 2× (no control) could be due to genetic variability from segregating population for the glyphosate resistance. The lack of control at the label rate is consistent with the survival of A. palmeri in the glyphosate-treated field from which its seeds were originally collected. The inclusion of glyphosate in glyphosate-tolerant maize, soybean, and cotton cultivation systems plays an important role in weed control in the RSA and accelerated the popularity of conservation tillage (Gouse Reference Gouse2014). Therefore, it is critical to manage weeds with glyphosate-resistance issues with the aim to preserve the continued benefits of glyphosate for weed control as well as for the other benefits of reduced-tillage practices, including maintenance of soil organic matter and beneficial insects, increased soil water-holding capacity, reduced soil and nutrient loss from the field, reduced soil compaction, and less time and labor required to prepare the field for planting (Duke and Powles Reference Duke and Powles2008; Kudsk and Mathiassen Reference Kudsk and Mathiassen2020). Resistance to glyphosate in A. palmeri is widespread and well documented (Dominguez-Valenzuela et al. Reference Dominguez-Valenzuela, Gherekhloo, Fernández-Moreno, Cruz-Hipolito, Alcántara-de la Cruz, Sánchez-Gonzáles and De Prado2017; Heap Reference Heap2022; Kaundun et al. Reference Kaundun, Jackson, Hutchings, Galloway, Marchegiani, Howell, Carlin, McIndoe, Tuesca and Moreno2019; Küpper et al. Reference Küpper, Borgato, Patterson, Netto, Nicolai, de Carvalho and Christoffoleti2017; Norsworthy et al. Reference Norsworthy, Griffith, Scott, Smith and Oliver2008; Steckel et al. Reference Steckel, Main and Ellis2008).
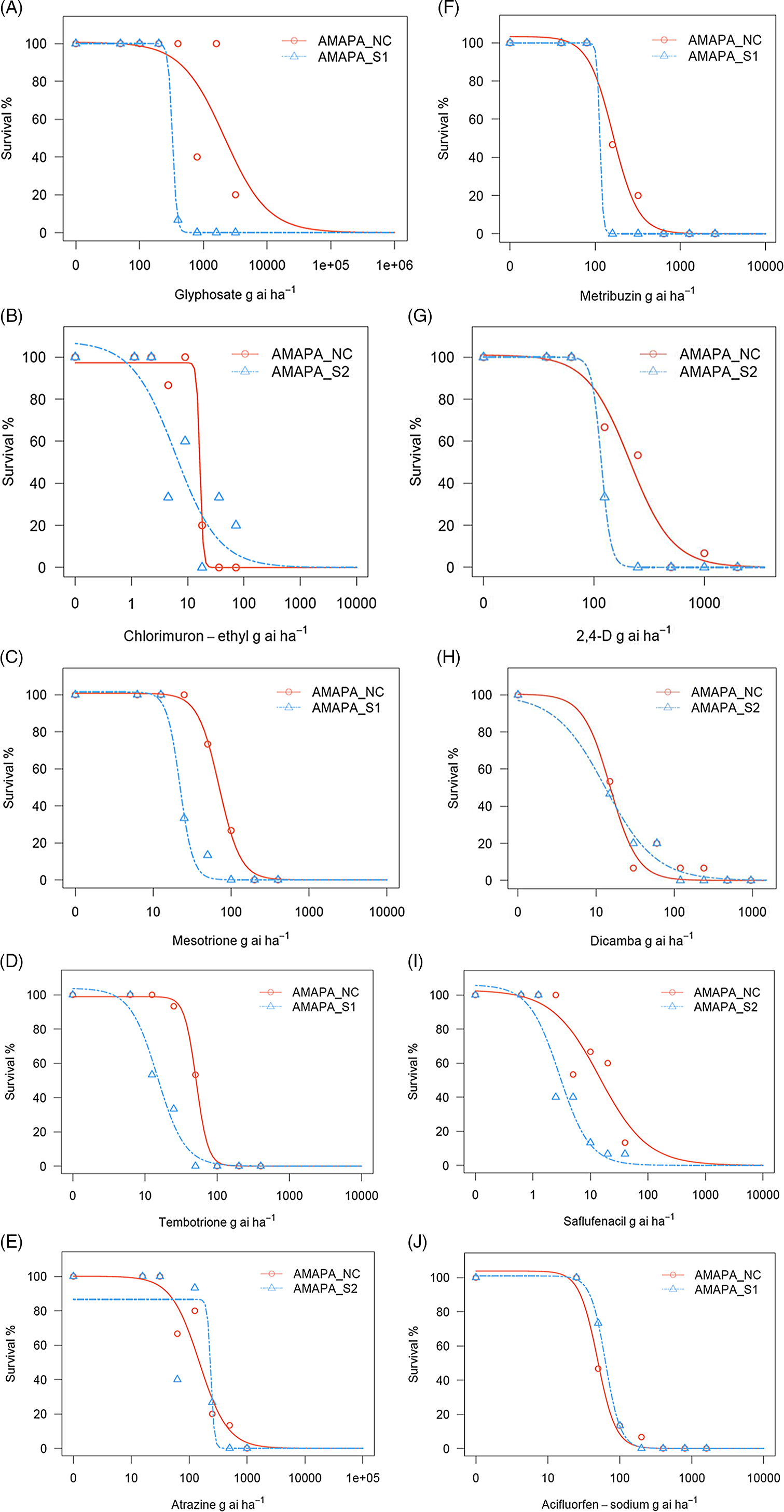
Figure 2. Percent survivors at various dose levels of postemergence herbicides including glyphosate (A), chlorimuron-ethyl (B), mesotrione (C), tembotrione (D), atrazine (E), metribuzin (F), 2,4-D (G), dicamba (H), saflufenacil (I), and acifluorfen-sodium (J) applied to the Amaranthus palmeri population AMAPA-NC, found in the Republic of South Africa, and to two sensitive populations, AMAPA-S1 (Shickley, NE, USA) and AMAPA-S2 (Macon, GA, USA). Each herbicide treatment included seven different rates. Visual survival rating was assessed at 16 d after treatment. Dose–response analysis was conducted using a three-parameter log-logistic model using the drc package in R software per Seefeldt et al. (Reference Seefeldt, Jensen and Fuerst1995).
The efficacy of chlorimuron-ethyl, an ALS-inhibiting herbicide (Group 2), showed variation ranging from 100% control at both 2× and 4× the label rate and 80% control at the label rate (18 g ae ha−1), to little or no control at all of the sublethal rates tested (Table 4; Figure 2B). Resistance to ALS inhibitors in A. palmeri populations is widely reported (Burgos et al. Reference Burgos, Kuk and Talbert2001; Heap Reference Heap2022; Singh et al. Reference Singh, Singh, Salas-Perez, Bagavathiannan, Lawton-Rauh and Roma-Burgos2019; Wise et al. Reference Wise, Grey, Prostko, Vencill and Webster2009), and as of January 2022, there were 167 cases of weed species documented to be resistant to ALS inhibitors globally (Heap Reference Heap2022). Herbicidal efficacy similar to that of chlorimuron-ethyl (70% to 80% control at the label rate) was observed for mesotrione (HPPD inhibitor; Group 27) and atrazine (PS II inhibitor; Group 5), with the exception that atrazine also showed reduced control (87%) at 2× the label rate in the AMAPA-NC population (Table 4; Figure 2C and E). Interestingly, metribuzin (another Group 5 herbicide) was significantly more effective than atrazine (Table 4; Figure 2F), indicating the possibility of NTSR mechanisms such as reduced absorption and/or translocation, increased sequestration, and enhanced metabolic degradation (Rigon et al. Reference Rigon, Gaines, Küpper and Dayan2020). Among these NTSR mechanisms, metabolism-based herbicide resistance represents a major threat, because it can impart resistance to herbicides from varied chemical classes across several mechanisms of action (Rigon et al. Reference Rigon, Gaines, Küpper and Dayan2020; Yu and Powles Reference Yu and Powles2014). The large and ubiquitous family of cytochrome P450 monooxygenase (P450) enzymes are implicated in increased herbicide metabolism as a mechanism of resistance in certain weed species (Cobb and Reade Reference Cobb and Reade2010; Katagi and Mikami Reference Katagi, Mikami and Roberts2000). GSTs are a superfamily of enzymes that catalyze the conjugation of various substances, including herbicides, to the tripeptide glutathione (GSH) (Cobb and Reade Reference Cobb and Reade2010). GSTs have been implicated in resistance to atrazine in velvetleaf (Abutilon theophrasti Medik.) (Devine and Preston Reference Devine and Preston2000) and A. tuberculatus (Ma et al. Reference Ma, Kaundun, Tranel, Riggins, McGinness, Hager, Hawkes, McIndoe and Riechers2013), and the P450 system confers resistance to metribuzin and simazine in rigid ryegrass (Lolium rigidum Gaudin). Anderson and Gronwald (Reference Anderson and Gronwald1991), Gray et al. (Reference Gray, Stoltenberg and Balke1996), and Nakka et al. (Reference Nakka, Godar, Thompson, Peterson and Jugulam2017a) also suggested a role for GSTs in the metabolism of atrazine. GST-mediated metabolism reportedly also contributes to trifluralin resistance in A. palmeri (González-Torralva and Norsworthy Reference González-Torralva and Norsworthy2021; Gossett et al. Reference Gossett, Murdock and Toler1992).
Interestingly, another Group 27 herbicide, tembotrione, provided full control at the label rate or higher, indicating differential efficacies between mesotrione and tembotrione; this is in contrast to a report that showed cross-resistance between these herbicides belonging to the same triketone chemical family in an A. tuberculatus population (Kaundun et al. Reference Kaundun, Hutchings, Dale, Howell, Morris, Kramer, Shivrain and Mcindoe2017). However, reduction in sensitivity was also seen for tembotrione at sublethal rates in the AMAPA-NC population (Table 4; Figure 2D). Increased target-site gene expression and higher HPPD protein levels have been shown to contribute to mesotrione resistance in A. palmeri (Betha et al. Reference Betha, Thompson, Peterson and Jugulam2017; Nakka et al. Reference Nakka, Godar, Wani, Thompson, Peterson, Roelofs and Jugulam2017b); however, these data included a limited number of individual plants with high variability (Kaundun et al. Reference Kaundun, Hutchings, Dale, Howell, Morris, Kramer, Shivrain and Mcindoe2017). Metabolic resistance via rapid detoxification of mesotrione, coupled with increased HPPD gene expression, has been reported in A. palmeri (Nakka et al. Reference Nakka, Godar, Wani, Thompson, Peterson, Roelofs and Jugulam2017b; Thompson et al. Reference Thompson, Peterson and Lally2012), and these mechanisms were also found to be involved in cross-resistance to HPPD-inhibiting herbicides in a R. raphanistrum population in Australia that is also resistant to atrazine, chlorsulfuron, diflufenican, and 2,4-D (Lu et al. Reference Lu, Yu, Han, Owen and Powles2020a). Metabolic resistance through the activity of cytochrome P450 enzymes is implicated in tembotrione metabolism in A. palmeri (Küpper et al. Reference Küpper, Peter, Zöllner, Lorentz, Tranel, Beffa and Gaines2018b). Cytochrome P450 involvement in rapid detoxification resulting in limited translocation of mesotrione has been demonstrated as a resistance mechanism in an A. tuberculatus biotype in Nebraska, USA (Kaundun et al. Reference Kaundun, Hutchings, Dale, Howell, Morris, Kramer, Shivrain and Mcindoe2017) and another A. tuberculatus biotype in Illinois, USA (Ma et al. Reference Ma, Kaundun, Tranel, Riggins, McGinness, Hager, Hawkes, McIndoe and Riechers2013).
The greatest reduction in A. palmeri control among the herbicides tested was observed for saflufenacil (PPO inhibitor; Group 14) (Table 4; Figure 2I). It showed less than 50% control at the 2×, 1×, and 1/2× rates and no control at any of the other sublethal doses tested, indicating potential resistance to saflufenacil. However, acifluorfen, a Group 14 herbicide that is not yet registered in the RSA, showed excellent control of the AMAPA-NC population (Table 4; Figure 2J). The differential response between saflufenacil and acifluorfen could be due to differences in their chemical family (N-phenyl-amides and diphenyl ethers, respectively) and/or due to no history of exposure to acifluorfen, despite both being Group 14 herbicides (HRAC 2021). Resistance to both saflufenacil and acifluorfen has been reported in wild poinsettia (Euphorbia heterophylla L.) in Brazil (Heap Reference Heap2022).
Many of the postemergence herbicides were effective in controlling the AMAPA-NC population (Table 4). Glufosinate (Group 10) is a nonselective, fast-acting postemergence herbicide and was shown to be highly effective at various application rates, including 1/2× the label rate, to control the AMAPA-NC population (Table 4). However, the use of glufosinate as a postemergence, in-crop treatment to manage weeds, including A. palmeri, would require glufosinate-tolerant crops (Hoffner et al. Reference Hoffner, Jordan, Chandi, York, Dunphy and Everman2012). Recently, a biotype of A. palmeri was reported to be resistant to glufosinate in the United States (Heap Reference Heap2022). Previously, resistance to glufosinate had been reported in biotypes of Lolium spp. and goosegrass [Eleusine indica (L.) Gaertn.] grass weeds (Heap Reference Heap2022). Resistance to glufosinate in Lolium spp. was shown to be due to a target-site mutation at the plastidic glutamate synthase 2 (GS2) gene resulting in an aspartic acid to asparagine change at amino acid position 171 (Avila-Garcia et al. Reference Avila-Garcia, Sanchez-Olguin, Hulting and Mallory-Smith2012). However, the exact mechanism(s) of resistance to glufosinate in E. indica remains unknown (Jalaludin et al. Reference Jalaludin, Yu, Zoellner, Beffa and Powles2017).
Auxin mimic (Group 4) herbicides, including 2,4-D and dicamba, which belong to the phenoxy-carboxylates and benzoates, respectively, exhibited good control at and above the label rates (Table 4; Figure 2G and H). Dicamba showed >90% control at 1/2× the label rate, while reduction in sensitivity was observed for 2,4-D at 1/2× the label rate (Table 4; Figure 2G). A total of nine broadleaf weed species worldwide have been confirmed to be resistant to dicamba to date—a population of A. palmeri, prickly lettuce (Lactuca serriola L.), and several populations of B. scoparia in the United States; a population of R. raphanistrum in Australia; lambsquarters (Chenopodium album L.) in New Zealand; common hempnettle (Galeopsis tetrahit L.), B. scoparia, and wild mustard (Sinapis arvensis L.) in Canada; A. hybridus in Argentina; S. arvensis in Turkey; and bachelor’s button/cornflower (Centaurea cyanus L.) in Poland (Heap Reference Heap2022). For 2,4-D, a total of 26 broadleaf weeds were reported to be resistant worldwide, including some of the Amaranthus species (A. palmeri and A. tuberculatus in the United States; A. hybridus in Argentina) (Heap Reference Heap2022; Kumar et al. Reference Kumar, Liu, Boyer and Stahlman2019). Mutations that reduce the binding of synthetic auxin herbicides to a receptor protein (auxin-binding proteins [auxin F-box proteins or AFB]) and to a co-receptor protein (indole-3-acetic acid or Aux/IAA protein) have been shown to confer resistance in weeds, as in the case of S. arvensis (Gaines et al. Reference Gaines, Duke, Morran, Rigon, Tranel, Küpper and Dayan2020; Webb and Hall Reference Webb and Hall1995). In a recent study, it was demonstrated that a single amino acid change in IAA16 protein in B. scoparia could result in cross-resistance to dicamba, 2,4-D, and fluroxypyr herbicides (LeClere et al. Reference LeClere, Wu, Westra and Sammons2018). Weed control by use of herbicides within integrated weed management programs must meet a minimum control standard (80% for direct field treatments at the recommended rate) to be accepted by farmers (Frans et al. Reference Frans, Talbert, Marx and Crowley1986; Nunes et al. Reference Nunes, Vidal, Trezzi, Kalsing and Goulart2007). Overall, our results indicated that tembotrione (Group 27), metribuzin (Group 5), acifluorfen (Group 4), 2,4-D and dicamba (Group 4), and glufosinate (Group 10) are highly effective postemergence herbicides to be considered in a weed management program to control the AMAPA-NC population, while unsatisfactory control can be expected with single-SOA treatments applied postemergence with glyphosate (Group 9), chlorimuron-ethyl (group 2), mesotrione (Group 27), atrazine (Group 5), and saflufenacil (Group 14).
It is generally accepted that SOA diversity in herbicide programs is a key driver for successful resistance management (Beckie Reference Beckie2020; Busi et al. Reference Busi, Powles, Beckie and Renton2020; Comont et al. Reference Comont, Lowe, Hull, Crook, Hicks, Onkokesung, Beffa, Childs, Edwards, Freckleton and Neve2020; Evans and Joshi Reference Evans and Joshi2016). While the current study was focused on understanding sensitivities to key herbicides individually, it is important to diversify weed management programs with effective herbicides with different SOAs used in mixtures or in rotation as key tools in resistance management strategies. The herbicide sensitivity profiling of the AMAPA-NC Palmer amaranth population from our study was broadly shared in the RSA via industry groups, universities, and government agencies to create awareness (CropLife South Africa 2019) as well as to support further on-farm research to explore combination of SOAs for effective control. Mixtures of effective herbicides with different SOAs do not prevent the evolution of resistance but have the potential to delay it relative to repeated use of a single herbicide. In addition, herbicide mixtures will also suppress weed populations that are already resistant to herbicides with some SOAs, thereby further delaying the economic impacts of resistance (Busi et al. Reference Busi, Powles, Beckie and Renton2020; Comont et al. Reference Comont, Lowe, Hull, Crook, Hicks, Onkokesung, Beffa, Childs, Edwards, Freckleton and Neve2020; Diggle et al. Reference Diggle, Neve and Smith2003; Evans et al. Reference Evans, Tranel, Hager, Schutte, Wu, Chatham and Davis2016; Liu et al. Reference Liu, Neve, Glasgow, Wuerffel, Owen and Kaundun2020).
Molecular Characterization of Potential Target-Site Resistance
To identify potential molecular mechanisms for resistance and/or reduced sensitivity to certain herbicides observed in the AMAPA-NC population (Tables 3 and 4), we carried out gene sequencing of target-site mutations that were previously shown to confer resistance to glyphosate (EPSPS), chlorimuron-ethyl (ALS), atrazine (PS II), and saflufenacil (PPO).
No Known Target-Site Mutations Were Found in EPSPS
EPSPS gene amplification through PCR from 20 individual plants of the AMAPA-NC population produced an expected 195-bp fragment encompassing EPSPS codons 102, 103, and 106. Gene sequencing of the PCR-amplified EPSPS nucleotide sequences showed 100% homology with previously published A. palmeri data (e.g., GenBank references: ACV53021.1 and ACV53022.1), confirming the identity of the EPSPS fragment. The EPSPS sequence comparison between AMAPA-NC and the susceptible reference sequences (Figure 3) showed none of the previously reported mutations in codons 102, 103, or 106 (Li et al. Reference Li, Peng, Han, Nyporko, Kulynych, Yu and Powles2018; Perotti et al. Reference Perotti, Larran, Palmieri, Martinatto, Alvarez, Tuesca and Permingeat2019; Sammons and Gaines Reference Sammons and Gaines2014; Yu et al. Reference Yu, Jalaludin, Han, Chen, Sammons and Powles2015), indicating that the resistance to glyphosate observed in this population (Table 4; Figure 2A) is not likely due to the known target-site mutations in the EPSPS gene.

Figure 3. Translated protein sequence of the Amaranthus palmeri EPSPS gene sequence. Thr102, Ala103, and Pro106 are highlighted in red, showing no target-site mutations in the plants investigated.
Elevated Levels of EPSPS Gene Copy Number
To further dissect the potential mechanism of resistance to glyphosate, we used quantitative RT-PCR to measure relative genomic copy numbers of the EPSPS gene relative to the ALS gene (single-copy control) in 20 individual AMAPA-NC plants. Amaranthus hybridus plants collected at the same site were used as susceptible control plants with one EPSPS copy. Genomic EPSPS copy numbers relative to ALS copy number ranged from 2 to 140 (decimals rounded off to the nearest integer) in the AMAPA-NC population, with an average gene copy number of 75 (Figure 4), while the susceptible A. hybridus plants (“AMAHY”) had a relative copy number of 1. High EPSPS gene copy number is considered to be a mechanism of resistance to glyphosate in A. palmeri (Gaines et al. Reference Gaines, Zhang, Wang, Bukun, Chisholm and Shaner2010) and other weed species, including monocots and dicots (Patterson et al. Reference Patterson, Pattinga, Ravet, Neve and Gaines2018; Powles Reference Powles2010). High EPSPS copy number in A. palmeri, a diploid weed species (Rayburn et al. Reference Rayburn, McCloskey, Tatum, Bollero, Jeschke and Tranel2005), has been shown to be due to extrachromosomal circular DNA containing the EPSPS gene cassette (Koo et al. Reference Koo, Molin, Saski, Jiang, Putta, Jugulam, Friebe and Gill2018b). Our result confirmed that the observed resistance to glyphosate in the AMAPA-NC population (Table 4) is mainly conferred by high EPSPS gene copy number. Because glyphosate bound to the EPSPS protein becomes unavailable, an increase in the EPSPS protein pool will require an equivalent increase in the glyphosate dose needed to cause plant death (Gaines et al. Reference Gaines, Zhang, Wang, Bukun, Chisholm and Shaner2010). Previous studies of glyphosate-resistant A. palmeri have reported EPSPS copy numbers between 30 and 179 (Gaines et al. Reference Gaines, Zhang, Wang, Bukun, Chisholm and Shaner2010; Küpper et al. Reference Küpper, Borgato, Patterson, Netto, Nicolai, de Carvalho and Christoffoleti2017). Küpper et al. (Reference Küpper, Borgato, Patterson, Netto, Nicolai, de Carvalho and Christoffoleti2017) further showed lower levels of accumulated shikimic acid and higher levels of resistance associated with higher EPSPS copy number.
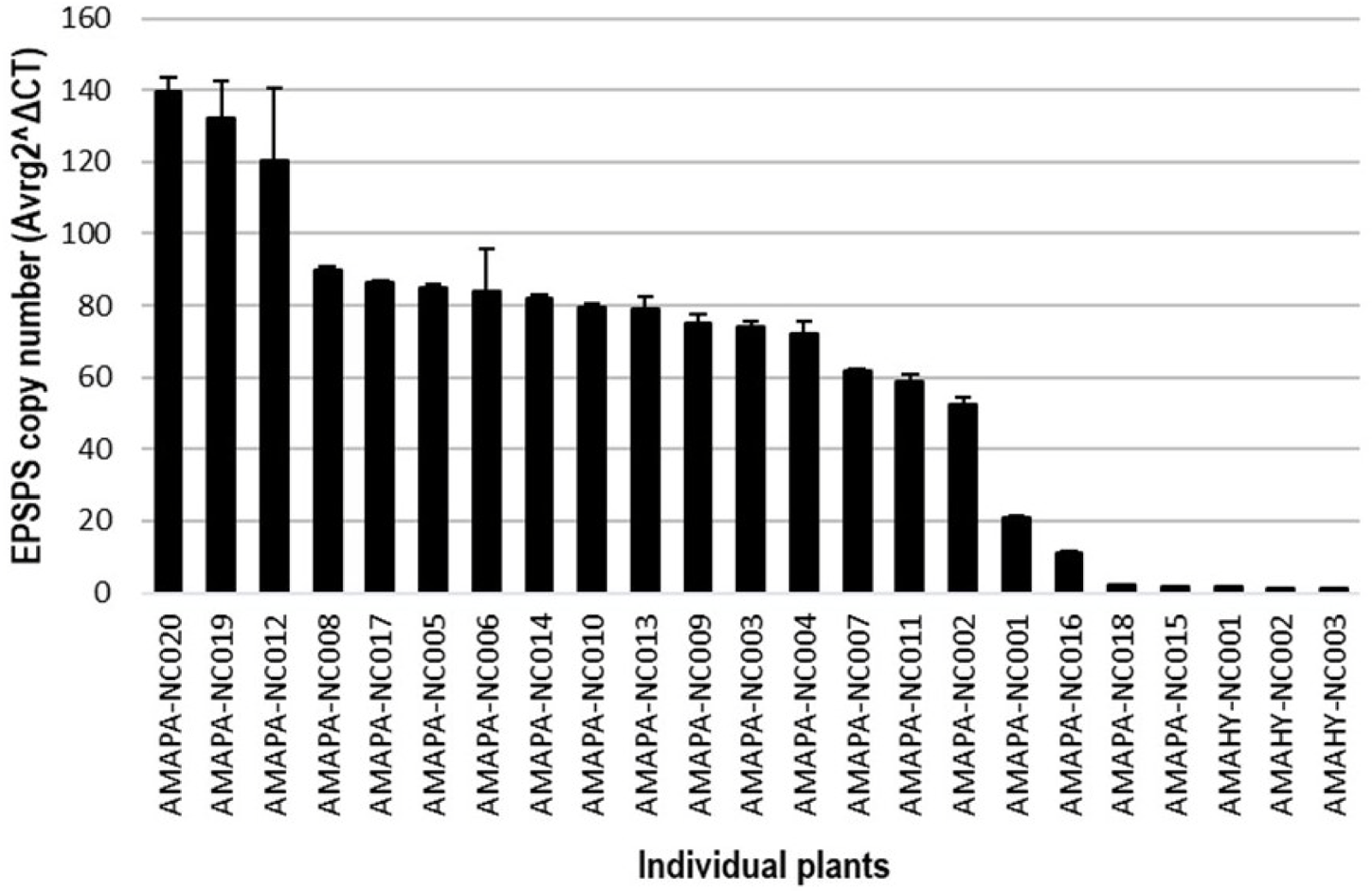
Figure 4. EPSPS gene copy number in Amaranthus palmeri plants of the AMAPA-NC population and in glyphosate-susceptible Amaranthus hybridus (AMAHY) plants collected at the same site.
Known Ser-653-Asn Target-Site Mutation Found in the ALS Gene
Resistance-conferring mutations in the ALS enzyme occur primarily in two domains: CAD, comprising amino acids 124 to 205 (based on Arabidopsis sequence as a standard); and BE, comprising amino acids 574 to 653 (Shaner Reference Shaner1991; Socorro Reference Socorro2011). PCR amplification of ALS genes from 20 individual plants of the AMAPA-NC A. palmeri population resulted in the expected 420-bp fragment for the CAD domain (including target codons 122, 197, and 205) and the expected 340-bp fragment for the BE domain, which included codons 574 and 653. The ALS predicted protein sequence comparison between AMAPA-NC and the susceptible reference sequence of A. palmeri (GenBank accession: KT833339.1), a susceptible A. hybridus sequence (AMAHY-NC001), and a resistant A. palmeri reference sequence (GenBank accession: ASL69937.1) showed no mutations in the CAD domain (data not shown). In the BE domain, no mutations were found at position 574, but for position 653, nearly half (8/20) of the tested plants contained the Ser-653-Asn mutation (Figure 5). This mutation is associated with high levels of resistance to imidazolines and pyrimidinylthiobenzoates and with low levels of resistance to sulfonylureas (Berger et al. Reference Berger, Madeira, Ferrell, Gettys, Morichetti, Cantero and Nunez2016; Patzoldt and Tranel Reference Patzoldt and Tranel2007; Saari et al. Reference Saari, Cotterman and Thill1994; Tranel and Wright Reference Tranel and Wright2002), indicating that the resistance to chlorimuron-ethyl observed in this population (Table 4; Figure 2B) is likely due to the Ser-653-Asn target-site mutation in the ALS gene.

Figure 5. Translated protein alignment of the ALS BE domain showing no mutations at position 574 (W) and a subset of plants with mutations at position 653 (Ser-653-Asn).
No Known Target-Site Mutations Found in PPX2 Gene in Codon 98 or 210
Based on the dCAPS assay (Giacomini et al. Reference Giacomini, Umphres, Nie, Mueller, Steckel, Young, Scott and Tranel2017), no known point mutations were found in codon 98 (also referred as codon 128) of the PPX2 gene in any of the 19 individual AMAPA-NC plants analyzed (Figure 6). In addition, sequence analysis of the region containing codon 210 did not show any known mutation of the glycine residue (Salas et al. Reference Salas, Burgos, Tranel, Singh, Glasgow, Scott and Nichols2016). This indicates that the observed reduced sensitivity to saflufenacil in this population (Table 4; Figure 2I) is not likely due to known target-site mutations in the PPX2 gene. A recently identified PPX2 gene mutation at position 399 with a glycine to alanine substitution (G399A) in A. palmeri (Rangani et al. Reference Rangani, Salas-Perez, Aponte, Knapp, Craig, Meitzner, Langaro, Noguera, Porri and Roma-Burgos2019) is unlikely to be the cause for saflufenacil resistance in the AMAPA-NC population; first, because the G399A mutation is shown to confer resistance to broad-spectrum PPO inhibitors, including acifluorfen and saflufenacil herbicides. In contrast, we found acifluorfen to be highly effective (100% and 93% control at the label and 1/2× the label rate, respectively; Table 4) compared with the significantly reduced efficacy of saflufenacil (33% control at the label rate). Second, the presence of a G399A mutation in A. palmeri population has not always been shown to confer resistance to saflufenacil (Wu et al. Reference Wu, Goldsmith, Pawlak, Feng, Smith, Navarro and Perez-Jones2020). Further evaluation is needed to understand the potential cause of resistance to saflufenacil herbicide in the AMAPA-NC A. palmeri population by either analyzing any hitherto unknown target-site mutation/s in the PPX2 gene and/or exploring the role of NTSR for any differential detoxification/metabolism of saflufenacil compared with acifluorfen.
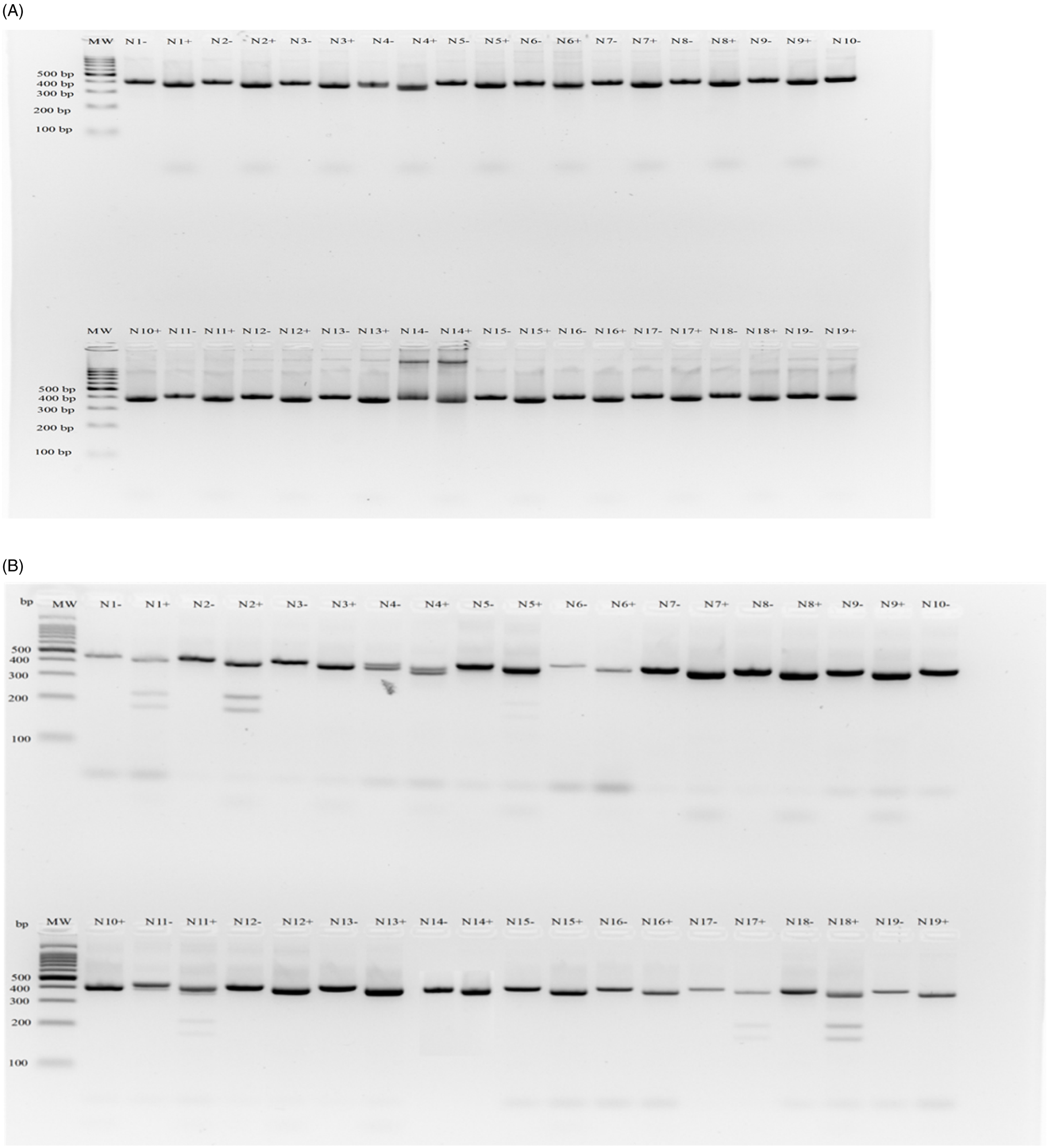
Figure 6. Gel image of dCAPS assay for AMAPA-NC plants collected in the Northern Cape. Accession labels were shortened for clear presentation of results (N1 = AMAPA-NC001, etc.). Both undigested (−) and digested (+) PCR products are shown. (A) Arg-98-Met assay; (B) Arg-98-Gly assay. A 100-bp DNA ladder (MW) (Thermo Fischer Scientific, Johannesburg, RSA) was used, and the PCR products fall between ∼400- and 350-bp fragments.
In conclusion, this investigation of a nonnative A. palmeri population (AMAPA-NC) in the RSA indicated resistance to chlorimuron-ethyl and glyphosate in this population, while decreased sensitivity was observed at the label rate for mesotrione, atrazine, saflufenacil, and S-metolachlor, and provided insight on mechanisms of resistance to selected herbicides. Moreover, this study provided detailed profiling of various preemergence and postemergence herbicides representing various SOAs that may serve as effective herbicide options in the design of an effective weed management program. Among the preemergence herbicides, acetochlor, isoxaflutole, diflufenican, and pyroxasulfone were found to be effective at controlling the A. palmeri AMAPA-NC population, and glufosinate, tembotrione, acifluorfen, dicamba, 2,4-D and metribuzin were effective as postemergence herbicide treatments.
Acknowledgments
We thank Graham Head and Aruna Varanasi for their useful comments and critical review of the draft article. CR and JV thank Bayer AG for financial support for the research conducted at the University of Pretoria as part of the South African Herbicide Resistance Initiative. The authors declare no conflict of interest.













