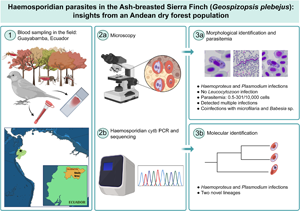
Introduction
Haemosporidian genera Plasmodium, Haemoproteus and Leucocytozoon, responsible for avian malarial infections, are highly diverse and infect birds in all continents but Antarctica (Valkiūnas, Reference Valkiūnas2005). They are obligate parasites transmitted by haematophagous dipteran vectors that infect blood cells and other host tissues (Valkiūnas, Reference Valkiūnas2005; Santiago-Alarcon et al., Reference Santiago-Alarcon, Palinauskas and Schaefer2012). The effects of avian malaria may range from mortality (e.g. Atkinson et al., Reference Atkinson, Woods, Dusek, Sileo and Iko1995; Palinauskas et al., Reference Palinauskas, Žiegytė, Iezhova, Ilgūnas, Bernotienė and Valkiūnas2016) to non-significant reductions in fitness (e.g. Bensch et al., Reference Bensch, Waldenström, Jonzán, Westerdahl, Hansson, Sejberg and Hasselquist2007; Hammers et al., Reference Hammers, Komdeur, Kingma, Hutchings, Fairfield, Gilroy and Richardson2016). During the acute, initial phase of the infection, birds may reduce food intake and body weight (Atkinson et al., Reference Atkinson, Dusek, Woods and Iko2000; Valkiūnas et al., Reference Valkiūnas, Z̆ic̆kus, Shapoval and Iezhova2006). They also increase their production of red blood cells (i.e. polychromasia; Palinauskas et al., Reference Palinauskas, Žiegytė, Šengaut and Bernotienė2022) to counteract the anaemia caused by the parasites (Mitchell and Johns, Reference Mitchell and Johns2008). Chronic infections are generally associated with milder symptoms (Goswami and Swamy, Reference Goswami and Swamy2013) but can generate trade-offs between immunological response and reproduction investment (e.g. Nordling et al., Reference Nordling, Andersson, Zohari and Lars1998; Asghar et al., Reference Asghar, Hasselquist and Bensch2011), and compromise long-term survival (e.g. Asghar et al., Reference Asghar, Palinauskas, Zaghdoudi-Allan, Valkiūnas, Mukhin, Platonova, Färnert, Bensch and Hasselquist2016).
The course of malarial infection is tightly related to the parasite genus (Atkinson and van Riper, Reference Atkinson, van Riper, Loye and Zuk1991). In general, Plasmodium is more pathogenic than Haemoproteus (Atkinson and van Riper, Reference Atkinson, van Riper, Loye and Zuk1991; Valkiūnas, Reference Valkiūnas2005), although highly pathogenic species of Haemoproteus have been reported (e.g. Sol et al., Reference Sol, Jovani and Torres2003). The pathogenicity of some species of Leucocytozoon can be high in domestic poultry and waterfowl, but its impact on other families of birds is poorly known (Atkinson and van Riper, Reference Atkinson, van Riper, Loye and Zuk1991). Also, infections with 2 or more species of haemosporidians are poorly understood (Marzal et al., Reference Marzal, Bensch, Reviriego, Balbontin and De Lope2008). Still, in a clinical trial comparing single and mixed infections, Palinauskas et al. (Reference Palinauskas, Žiegytė, Šengaut and Bernotienė2018) found that Plasmodium elongatum intensity of parasitaemia is enhanced by the presence of Plasmodium relictum. Also, in a field-based study, double infections caused a significant decline in body condition compared to single infections (Marzal et al., Reference Marzal, Bensch, Reviriego, Balbontin and De Lope2008).
Predicting the factors that drive avian malaria epidemiology is challenging because infections respond to a complex interplay between environmental and intrinsic factors. Water availability, in terms of permanent water sources or increased precipitation, tends to predict malaria prevalence (Okanga et al., Reference Okanga, Cumming and Hockey2013; Ferraguti et al., Reference Ferraguti, Martínez-de la Puente, Bensch, Roiz, Ruiz, Viana, Soriguer and Figuerola2018), because it limits vector reproduction and parasite development (Van Riper et al., Reference Van Riper, Van Riper, Goff and Laird1986; LaPointe et al., Reference Lapointe, Atkinson and Samuel2012). At the same time, host age, sex and body and health conditions can be intrinsic predictors of infection status and intensity of infection. For example, parasite prevalence may decline (e.g. van Oers et al., Reference van Oers, Richardson, Sæther and Komdeur2010; Hammers et al., Reference Hammers, Komdeur, Kingma, Hutchings, Fairfield, Gilroy and Richardson2016) or increase (e.g. Cosgrove et al., Reference Cosgrove, Wood, Day and Sheldon2008; Knowles et al., Reference Knowles, Wood, Alves, Wilkin, Bensch and Sheldon2011; Fecchio et al., Reference Fecchio, Lima, Silveira, Ribas, Caparroz and Marini2015) with age, depending on the host. Also, differences in prevalence between sexes have been related to sexual dimorphism (e.g. Svensson-Coelho et al., Reference Svensson-Coelho, Blake, Loiselle, Penrose, Parker and Ricklefs2013). The expectation is that males from species with higher dimorphism should suffer from higher prevalence because of the immunosuppressive effect of maintaining secondary sexual characters (Hamilton and Zuk, Reference Hamilton and Zuk1982; Zuk, Reference Zuk1990). However, sex-specific behaviours may cause higher exposure to parasite vectors or generate energy trade-offs between reproduction and immune response (Korpimäki et al., Reference Korpimäki, Hakkarainen and Bennett1993; van Oers et al., Reference van Oers, Richardson, Sæther and Komdeur2010; Baillie et al., Reference Baillie, Gudex-Cross, Barraclough, Blanchard and Brunton2012), increasing the susceptibility of the host. Additionally, weight, body condition or health condition before or during avian malarial infection could exert higher physiological stress on the host and decrease its ability to clear or control the infection (e.g. Lochmiller et al., Reference Lochmiller, Vestey and Boren1993; da Silva Rodrigues et al., Reference da Silva Rodrigues, de Souza Penha, Miwa, Branco and Junior2021).
We studied avian malarial infections in a population of the ash-breasted Sierra finch, Geospizopsis plebejus. This species is a common dweller of the Andean ecosystems, distributed from Ecuador south to Argentina, along almost the entire elevational range of this mountain chain, from sea level to 4500 m a.s.l. (Campagna et al., Reference Campagna, Geale, Handford, Lijtmaer, Tubaro and Lougheed2011). It is common in open habitats, including those disturbed by human activities (Jaramillo, Reference Jaramillo2021). Sexes are dimorphic in plumage (Jaramillo, Reference Jaramillo2021) but not significantly different in morphological measurements (Llerena-Quiroz, Reference Llerena-Quiroz2018). Females incubate (Hughes, Reference Hughes1980; Pozo-Zamora, Reference Pozo-Zamora2014), but no additional information on each sex's contributions to reproduction is known. Related species such as plumbeous Sierra finch, Geospizopsis unicolor and mourning Sierra finch, Rhopospina fruticeti, with similar latitudinal and elevational wide ranges (Campagna et al., Reference Campagna, Geale, Handford, Lijtmaer, Tubaro and Lougheed2011), have been studied as part of broad haemosporidian surveys (Merino et al., Reference Merino, Moreno, Vásquez, Martínez, Sánchez-Monsálvez, Estades, Ippi, Sabat, Rozzi and Mcgehee2008; Doussang et al., Reference Doussang, Sallaberry-Pincheira, Cabanne, Lijtmaer, González-Acuña and Vianna2021; McNew et al., Reference McNew, Barrow, Williamson, Galen, Skeen, DuBay, Gaffney, Johnson, Bautista, Ordoñez, Schmitt, Smiley, Valqui, Bates, Hackett and Witt2021). Still, G. plebejus has not received the same level of attention.
Here we present, for the first time, detailed information on the identity, prevalence and parasitaemia of haemosporidians and other haemoparasites that infect G. plebejus in an Andean dry forest. We also study the consequences of infection in the host's body and health condition and explore the environmental and intrinsic factors influencing infection status and parasitaemia. Given that, in general, Plasmodium tends to be more pathogenic than Haemoproteus (Valkiūnas, Reference Valkiūnas2005), we expected to find a higher prevalence and parasitaemia of Haemoproteus, because of its reduced fitness cost (Behomme et al., Reference Bedhomme, Agnew, Vital, Sidobre and Michalakis2005). Also, given the potential additive effect of double infections (e.g. Marzal et al., Reference Marzal, Bensch, Reviriego, Balbontin and De Lope2008; Palinauskas et al., Reference Palinauskas, Žiegytė, Šengaut and Bernotienė2018), we expected to find higher parasitaemia in individuals with coinfections. We also assumed that body and health conditions would be affected by infection status, parasite genus and coinfections. In terms of predictors, we expected that higher precipitation would favour higher prevalence and parasitaemia, because of increased possibilities for vector development (LaPointe et al., Reference Lapointe, Atkinson and Samuel2012). On the other hand, because the effect of age and sex on malarial infections tends to be host-specific, we had no clear expectations about the impact of these 2 variables. Finally, we expected that individuals with poor body and health conditions should have a higher probability of infection and higher parasitaemias than those with good body and health conditions (da Silva Rodrigues et al., Reference da Silva Rodrigues, de Souza Penha, Miwa, Branco and Junior2021).
Materials and methods
Study area and sampling
We sampled G. plebejus at Bosque Protector Jerusalem (BPJ) between December 2012 and June 2013, as part of a broader project focused on generating a community-level baseline for the avian host-parasite dynamics in this site. The BPJ is a public, protected area located in the Guayllabamba valley, 10 km north of Quito, in north-western Ecuador (00°00′17.4″N/078°21′34.7″W; 2000–2500 m a.s.l.). This area encompasses 1110 ha of protected inter-Andean dry forest remnants. It presents seasonality in water availability, with a wet season from October to May (average monthly precipitation for 2012–2013: 66.37 mm) and a dry season from June to September (average monthly precipitation for 2012–2013: 7.7 mm) (data provided by Instituto Nacional de Meteorología e Hidrología, INAMHI). The average temperature remains uniform throughout the year but varies markedly during the day. The average monthly temperature during the study period was 17.3°C, whereas the minimum and maximum average monthly temperatures were 11.8 and 25.2°C, respectively. The vegetation of BPJ is semi-deciduous scrubland of the northern valleys of the Andes (MAE, 2013). Xerophytic species such as Algarrobo (Acacia macracantha), yellow trumpetbush (Tecoma stans), tuna cactus (Opuntia soederstromiana) and quishuar (Buddleja bullata) are characteristic of this forest and dominate the landscape (Guerrón et al., Reference Guerrón, Orellana, Loor and Zambrano2005).
We set 4 sampling sites encompassing different microclimates, at least 300 m apart (Fig. 1). Sites 1 and 3 were in relatively undisturbed areas along a trail surrounded by native vegetation. Site 2 was located on a plantation of lime (Citrus sp.) and avocado (Persea americana), and site 4 was in a disturbed area of the forest, used for recreational purposes. Sites 1 and 3 were separated by 600 m from sites 2 and 4. We placed 7 mist nets at each site (dimensions: 12 m × 2.5 m, mesh: 36 mm, 4 shelves), targeting all the species that could get trapped in nets. We sampled each site in 1.5 day monthly visits for a yield of 6 samplings per site, separated by 4 weeks. Each sampling started at 6:00 and ended at 18:00 on the first day and at 12:00 on the second day. All individuals captured were sexed by plumage, measured (tarsus length, bill length, bill width, bill height and wing length) and ringed with plastic bands before release to track recaptures of individuals marked during this study. We drew blood samples from the jugular or brachial veins using syringes attached to 27G disposable needles. Blood smears were prepared in the field using 15–20 μL of blood and then fixed using 100% methanol for 3 min. In the laboratory, fixed blood smears were stained with 10% Giemsa solution for 1 h. The remaining blood samples were stored in 99% ethanol solution for molecular analysis.

Fig. 1. Map of the study area. Sampling sites (1–4) are shown in yellow inside BPJ (red contour).
Molecular lineage identification and phylogenetic inference
We extracted whole genomic DNA from blood samples using an in-house protocol (Peñafiel et al., Reference Peñafiel, Flores, Rivero De Aguilar, Guayasamin and Bonaccorso2019). We amplified partial sequences of the mitochondrial cytochrome b (cytb) gene of Plasmodium and Haemoproteus using the HaemF and HaemR2 primers (Bensch et al., Reference Bensch, Stjernman, Hasselquist, Örjan, Hannson, Westerdahl and Pinheiro2000). Amplification reaction solutions (25 μL volume) consisted of 5 μL of genomic DNA, 1× buffer, 3 mm MgCl2, 0.4 mm dNTPs, 0.6 μ m HaemF, 0.6 mm HaemR2 and 0.05 U μL−1 platinum Taq polymerase (Invitrogen Inc., Carlsbad, USA). We applied a non-nested polymerase chain reaction (PCR) protocol consisting of an initial 3 min denaturation at 94°C; 37 cycles of 94°C denaturation for 30 s, 50°C annealing for 30 s and 72°C extension for 45 s; and 75°C final extension for 10 s. We visualized amplicons by electrophoresis in a 1.2% agarose gel stained with SYBR Safe (Invitrogen Inc., Carlsbad, USA). Positive amplicons were treated with ExoSAP-IT (Affymetrix Inc., Santa Clara, USA) and Sanger sequenced on an ABI 3730XL sequence analyser (Applied Biosystems Inc., Foster City, USA) with the same PCR primers.
We assembled consensus sequences using Geneious 5.1.7 (Kearse et al., Reference Kearse, Moir, Wilson, Stones-Havas, Cheung, Sturrock, Buxton, Cooper, Markowitz, Duran, Thierer, Ashton, Meintjes and Drummond2012). All sequences showing no coinfections in chromatograms (i.e. double peaks) were aligned using Clustal X2.1 (Thompson, Reference Thompson1997). We used GenBank and MalAvi BLASTN tools (Zhang et al., Reference Zhang, Schwartz, Wagner and Miller2000; Bensch et al., Reference Bensch, Hellgren and Pérez-Tris2009) to compare our sequences with previously published ones. New lineages were those with less than 100% query match. We produced separate alignments for unique sequences of Plasmodium and Haemoproteus, using Clustal X2.1. We used Leucocytozoon fringillinarum as the outgroup in both alignments (GenBank accession number: JQ815435). We used Mesquite 3.61 (Maddison and Madison, Reference Maddison and Madison2019) to crop the alignment to match the outgroup sequence length (474 base pairs). We also translated the alignment into amino acids to verify the absence of stop codons or indels. Unphased sequences (i.e. showing potential coinfections) were analysed in DnaSP 6.12 (Rozas et al., Reference Rozas, Ferrer-Mata, Sánchez-DelBarrio, Guirao-Rico, Librado, Ramos-Onsins and Sánchez-Gracia2017) using the algorithm PHASE (Stephens et al., Reference Stephens, Smith and Donnelly2001; Stephens and Donnelly, Reference Stephens and Donnelly2003). We ran the program using the FASTA sequences of coinfections with International Union of Pure and Applied Chemistry ambiguity codes, including the unique sequences of the lineages found in the sample. Program settings for the analyses were: 1000 iterations, thinning interval = 10, burn-in = 200 and no recombination.
We used PartitionFinder 2.1.1 to select the best model of molecular evolution for each alignment (Guindon et al., Reference Guindon, Dufayard, Lefort, Anisimova, Hordijk and Gascuel2010; Lanfear et al., Reference Lanfear, Frandsen, Wright, Senfeld and Calcott2016). The parameters used were branch lengths linked, corrected Akaike information criterion (AICc), and greedy algorithm, an algorithm for heuristic search that increases the efficiency in finding the optimum partitioning scheme while reducing the number of schemes that need to be considered for a given dataset (Lanfear et al., Reference Lanfear, Calcott, Ho and Guindon2012). The best partition schemes for Plasmodium (by codon position from first to third) were: GTR (generalized time-reversible) + I (invariable sites) + G (gamma distribution); F81 (Felsenstein 4-parameter) and GTR + G, respectively. The best partition schemes for Haemoproteus (by codon position from first to third) were: GTR + I + G; GTR + G and GTR + G. We generated phylogenetic trees using MrBayes 3.2.7 for Bayesian inference (Ronquist and Huelsenbeck, Reference Ronquist and Huelsenbeck2003) and W-IQ-TREE 1.6.12 for maximum-likelihood inference (Trifinopoulos et al., Reference Trifinopoulos, Nguyen, von Haeseler and Minh2016). The analysis in MrBayes was set for 10 million generations, sampling every 1000 trees, discarding 2000 and retaining 8000 trees. The analysis in W-IQ-TREE was set for 10 000 bootstrap replicates using Ultrafast Bootstrap (Hoang et al., Reference Hoang, Chernomor, von Haeseler, Minh and Vinh2018). The resulting trees were edited with FigTree 1.4.3 (Rambaut, Reference Rambaut2017).
Morphological identification of haemoparasites
Blood smears were double-blind diagnosed and photographed at 1000× magnification using an Olympus BX43 microscope coupled with a DP27 camera and its software, CellSens (Olympus Corporation, Japan). We obtained standard morphometric measurements from uninfected erythrocytes, complex erythrocyte/parasites and parasites using ImageJ software (Schneider et al., Reference Schneider, Rasband and Eliceiri2012). Morphological identifications were based on keys from Valkiūnas (Reference Valkiūnas2005) and Valkiūnas and Iezhova (Reference Valkiūnas and Iezhova2018), as well as recent descriptions of new species (Walther et al., Reference Walther, Valkiūnas, González, Matta, Ricklefs, Cornel and Sehgal2014; Mantilla et al., Reference Mantilla, González, Lotta, Moens, Pacheco, Escalante, Valkiūnas, Moncada, Pérez-Tris and Matta2016).
Prevalence and parasitaemia
We considered a sample as infected if either PCR amplification or microscopic analysis diagnosed it as positive. Parasitaemia values (infected cells per 10 000 erythrocytes) were obtained from Giemsa-stained blood smear counts of positive individuals. Prevalence, mean parasitaemia and confidence intervals were calculated using Quantitative Parasitology 3.0 (Reiczigel et al., Reference Reiczigel, Marozzi, Fábián and Rózsa2019). Prevalence confidence intervals were calculated using the Sterne method (Reiczigel, Reference Reiczigel2003), and mean parasitaemia confidence intervals were obtained using the bias-corrected and accelerated (BCa) bootstrap interval method (Rózsa et al., Reference Rózsa, Reiczigel and Majoros2000). We calculated prevalence and mean parasitaemia for total haemosporidian infections (diagnosed by microscopy or PCR), genus (Haemoproteus and Plasmodium), parasite lineage and coinfections (detected by sequencing or morphological analysis). We excluded from these analyses all the samples from recaptures of individuals captured during the sampling period.
We compared the prevalence between parasites through chi-square tests, excluding coinfections. Also, after confirming that the parametric assumptions did not hold, we performed unpaired Wilcoxon rank-sum tests to compare the parasitaemia of individuals infected with different genera and those with single infections and coinfections. Finally, we also used Wilcoxon rank-sum tests to compare the body condition (residuals of the regression between body mass and tarsus length) and polychromatophil count per 10 000 erythrocytes examined between individuals infected and non-infected, infected with different parasites and bearing single infections and coinfections. High polychromatophil count, or polychromasia, indicates regenerative anaemia (Jones, Reference Jones2015), caused by various pathogens and health conditions (Mitchell and Johns, Reference Mitchell and Johns2008). Thus, it is a measure associated with the health status of the individual (e.g. Travers et al., Reference Travers, Clinchy, Zanette, Boonstra and Williams2010; Schoenle et al., Reference Schoenle, Kernbach, Haussmann, Bonier and Moore2017). Samples from recaptures were excluded from all analyses.
Predictors of infection status and parasitaemia
To explore how environmental and intrinsic factors may predict infection status and parasitaemia, we carried out a series of generalized linear model (GLM) analyses in R 1.2.5033 (R Development Core Team, 2013) using the MASS package (Venables and Ripley, Reference Venables and Ripley2002). We analysed infection status using a logit model (family = binomial, link = logit), where the response variable was the infection status obtained by PCR or microscopy, and analysed parasitaemia using a negative binomial regression model (link = log), where the response variable was the value of parasitaemia obtained through microscopy. The independent variables (predictors) used in both models were sex, age, body condition, polychromatophil count (as a measure of health condition), sampling site, average relative humidity of the month before the capture day and total precipitation of the month before the capture day. Environmental data were obtained from INAMHI. Totals and averages were calculated from daily values provided by INAMHI.
Model selection was performed as follows. Initially, we obtained a general model that included all factors. Then, we used the dropterm and stepaic functions (MASS package) to obtain 1-term deletions from the original complete model. All models were ranked based on the AICc, using the AICcmodavg package (Mazerolle, Reference Mazerolle2006) and the chosen (final) model was that with the highest AICc weight. Significance of the final model was assessed in MASS.
Results
We captured a total of 871 individuals from 36 bird species. Eighty-five birds were G. plebejus (9.76% relative abundance) and 65 of those were diagnosed by either molecular or microscopy methods. Out of these 65 individuals, 57 were infected with haemosporidians. Seven individuals were recaptured at least once during the sampling period (8.24%). Out of the recaptured individuals, 2 were captured at site 2 and recaptured at site 4, or vice versa, suggesting at least minimal exchange of individuals between these sites. Details for diagnoses by molecular and microscopic methods are provided in the following sections. Data for all the variables analysed are available in the Supplementary File 1: Dataset S1.
Molecular identification of haemosporidians
We applied PCR diagnosis to samples of 64 individuals, of which 52 were positive (Table 1). Among these positive samples, we identified 2 new Haemoproteus lineages: Haemoproteus GEPLE01 and GEPLE02 (GenBank accession numbers: ON938204 and ON938203, respectively). We also found 2 previously reported Haemoproteus sp. lineages: ZOCAP08 (originally called ZC1; GenBank: KC480265; Jones et al., Reference Jones, Cheviron and Carling2013) and AMAVIR01 (GenBank: JQ988544; McNew et al., Reference McNew, Barrow, Williamson, Galen, Skeen, DuBay, Gaffney, Johnson, Bautista, Ordoñez, Schmitt, Smiley, Valqui, Bates, Hackett and Witt2021). However, the phylogenetic position of these 2 lineages was unresolved within the major Haemoproteus lineages (Fig. 2). Additionally, samples showed infection by Plasmodium (Haemamoeba) cathemerium ZONCAP15 (GenBank: MK077679; Cadena-Ortiz et al., Reference Cadena-Ortiz, Mantilla, Rivero de Aguilar, Flores, Bahamonde, Matta and Bonaccorso2019; reported initially as ZOCAP15) and Plasmodium (Novyella) homopolare BAEBIC02 (GenBank: KF537287; González et al., Reference González, Lotta, García, Moncada and Matta2015). ZONCAP15 grouped with other P. cathemerium sequences, whereas BAEBIC02 grouped with P. homopolare sequences (Fig. 3).

Fig. 2. Phylogenetic position of the 4 lineages of Haemoproteus (in bold) found in the ash-breasted Sierra finch, Geospizopsis plebejus, among related Neotropical lineages. Bayesian posterior probabilities (Bpp) and maximum-likelihood bootstrap supports (MLb) are shown over nodes (Bbb/MLb). Each lineage includes: morphospecies or genus (if available); GenBank accession number (if available); MalAvi name and country where it was detected (CO, Colombia; CH, Chile; EC, Ecuador; ME, Mexico; PE, Peru; US, United States; NA, no information).

Fig. 3. Phylogenetic position of the 2 lineages of Plasmodium (in bold) found in the ash-breasted Sierra finch, G. plebejus, among related Neotropical lineages. Bpp and MLb are shown over nodes (Bbb/MLb). Each lineage includes: morphospecies or genus (if available); GenBank accession number (if available); MalAvi name (if available) and country where it was detected (BR, Brazil; CO, Colombia; CR, Costa Rica; CH, Chile; EC, Ecuador; PE, Peru; US, United States).
Table 1. Molecular lineages amplified by PCR and morphospecies detected in the ash-breasted Sierra finch, Geospizopsis plebejus, at BPJ, Ecuador

Four samples (HFC-321, HFC-425, HFC-718 and HFC-720) presented double peaks and were phased using DnaSP6 (Supplementary File 1: Coinfections), but only 2-phased sequences produced known lineages. Sample HFC-321 was infected with lineage AMAVIR01 (Haemoproteus sp.), which is the most common lineage found to be infecting the species in the study area (see Section ‘Prevalence and parasitaemia’), and a Haemoproteus sp. (as determined by BLASTN). Isolate HFC-720 produced lineage BAEBIC02 (P. homopolare) and Plasmodium sp. (as determined by BLASTN). The lineages in the other 2 isolates (HFC-425 and HFC-718) remained ambiguous; their phased sequences belonged to 2 Plasmodium sp. and 2 Haemoproteus sp., respectively.
Morphological identification and its correspondence with molecular lineages
We performed morphological identification on 65 individuals (Table 1), of which 51 were positive. Identified haemosporidians were as follows: Haemoproteus coatneyi, Haemoproteus erythrogravidus, P. homopolare (Fig. 4), 1 Plasmodium sp. and 3 new potential species of Haemoproteus. Additionally, we observed a coinfection of Haemoproteus sp. with Babesia sp. (isolate HFC-505) and another of H. coatneyi, Haemoproteus sp. and microfilaria (isolate HFC-677) (Fig. 4). We found no evidence of infection by Leucocytozoon.

Fig. 4. Haemoparasite stages observed in the ash-breasted Sierra finch, G. plebejus. (A) Microgametocyte, and (B) macrogametocyte of Haemoproteus coatneyi, (C) microgametocyte, and (D) macrogametocyte of Haemoproteus erythrogravidus, (E) erythrocytic meronts, (F) gametocyte of Plasmodium homopolare and (G, H) erythrocytic meronts of Babesia sp. Scale bar = 10 μm. (I, J) Confections of Plasmodium and Haemoproteus. Scale bar = 10 μm. (K, L) Microfilaria. Scale bar = 20 μm. Black arrowheads: pigment granules, double black arrowheads: merozoites. White arrowheads, protrusions of the erythrocyte membrane as the most relevant characteristics of H. erythrogravidus. Giemsa-stained thin blood smears. (A–H) At high magnification 1000×; (I–L) at low magnification 400×.
The correspondence between molecular lineages and morphospecies of haemosporidians is presented in Table 1. Seven out of 8 samples carrying lineages GEPLE01 or GEPLE02 were infected with different species of Haemoproteus. Out of 28 samples carrying lineage AMAVIR01, 14 were infected with H. coatneyi in single infections (10 samples) or coinfections (4 samples); the remaining samples showed H. erythrogravidus in single infection (7 samples) or coinfections (2 samples), or Haemoproteus sp. in single infections or coinfections. The only sample carrying lineage ZOCAP08 was associated with coinfection by H. erythrogravidus and an unidentified Haemoproteus species (Haemoproteus sp. 1). In the 6 samples carrying BAEBIC02, we identified P. homopolare and Plasmodium sp. in single infections or with H. erythrogravidus, and an unidentified Haemoproteus species. Only the molecular diagnosis detected infections by ZONCAP15. Also, we found mismatches between molecular and morphological diagnoses of coinfections (Table 1). None of the 19 coinfections detected by microscopy was detected by sequencing. Also, among the 4 samples with coinfections detected by sequencing, 2 were diagnosed as single infections of H. coatneyi by microscopy, and 2 had negative diagnoses.
Finally, we obtained diagnoses of the recapture(s) of 7 individuals (Table 2). Time lapses from capture to first or second recapture varied between <1 and 4 months. Unfortunately, the first capture of 1 individual (610 R), negative by microscopic diagnosis, could not be molecularly diagnosed. All the other individuals were infected in their first capture and in subsequent recaptures. Five of those were infected with the same lineage within <1 and 2 months.
Table 2. Molecular and morphological diagnosis of haemosporidian parasites in individuals of the ash-breasted Sierra finch, G. plebejus, captured and recaptured at BPJ, Ecuador

a Year, month, day.
Prevalence and parasitaemia
We found a high prevalence of haemosporidian parasite infections, with 87.7% of infected individuals (57 infected/65 analysed). Mean parasitaemia was 61.65 infected cells per 10 000 cells (N = 57) (Table 3). The prevalence of Haemoproteus molecular lineages was 56.9%, and their mean parasitaemia was 82.14 infected cells per 10 000 cells. The prevalence of Plasmodium lineages was 12.3%, and their mean parasitaemia was 22 infected cells per 10 000 cells. Prevalence and parasitaemia were higher for Haemoproteus lineages than for Plasmodium lineages (prevalence: chi-square = 28.58, df = 1, P < 0.0001; parasitaemia: W = 225, P = 0.023). The Haemoproteus sp. AMAVIR01 lineage showed the highest prevalence (43.1%) and highest mean parasitaemia (mean = 94.39 infected cells per 10 000 cells). The highest parasitaemia was found in HFC-695 sample (301 infected cells per 10 000 cells), which carried the AMAVIR01 lineage and H. erythrogravidus morphology. The only individual (HFC-535) infected with lineage ZOCAP08 (Haemoproteus sp.) presented high parasitaemia, with 219 infected cells per 10 000 cells. Microscopy of this sample revealed a coinfection between H. erythrogravidus and Haemoproteus sp. The prevalence of Babesia sp. and microfilariae was 1 in 65 individuals (1.54%).
Table 3. Prevalence and mean parasitaemia by haemosporidian parasites per parasite species and lineage the ash-breasted Sierra finch, G. plebejus, at BPJ, Ecuador

– denotes that no confidence interval is presented because of n = 1.
a Prevalence was calculated taking into account all positive samples by molecular and microscopic analysis excluding samples of recaptured individuals.
b Confidence interval. Prevalence confidence intervals were calculated using Sterne's method. Mean parasitaemia confidence intervals were calculated using the BCa bootstrap interval method.
c Parasitaemia was calculated from positive PCR samples, through microscopy (number of infected erythrocytes in 10 000 cells counted).
d Detected by molecular diagnosis or microscopy.
e Uncertain confidence intervals because of a small number of replicates.
We found no differences in body condition or polychromatophil count between positive and negative samples (body condition: W = 140, P = 0.13; polychromatophil count: W = 230, P = 0.98), samples carrying Haemoproteus and Plasmodium lineages (body condition: W = 142, P = 0.76; polychromatophil count: W = 127, P = 0.54), or samples carrying single infections and single coinfections (body condition: W = 276, P = 0.33; polychromatophil count: W = 368.5, P = 0.19). Parasitaemia was also not significantly different between samples carrying single infections and coinfections (W = 263.5; P = 0.059), but marginally significant results suggest that parasitaemia might be higher for coinfections if a larger sample was available.
Predictors of infection status and parasitaemia
In the analysis of predictors of infection status, model selection by AICc of the logit models retained a model with host age as the only predictor for infection status (AICcWt = 0.50; Table 4). According to this model, immature individuals show a lower prevalence than adults (Table 5). For parasitaemia, model selection by AICc of the negative binomial models retained the null model as the best model (AICcWt = 0.62; Table 6). This result precluded further exploration of the predictors of parasitaemia.
Table 4. Model selection criteria for the predictors of infection status by haemosporidian parasites (logit model) in individuals of the ash-breasted Sierra finch, G. plebejus, at BPJ, Ecuador

AICc, corrected Akaike information criterion; Cum., cumulative.
Table 5. Coefficient summary of the model for infection status by haemosporidian parasites (logit model) in individuals of the ash-breasted Sierra finch, G. plebejus, at BPJ, Ecuador

Table 6. Model selection criteria for the predictors of parasitaemia by haemosporidian parasites (negative binomial model) in the ash-breasted Sierra finch, G. plebejus, at BPJ, Ecuador

AICc, corrected Akaike information criterion; Cum., cumulative.
Discussion
Molecular and morphological diagnosis of haemosporidian parasites
We identified 6 cytb haemosporidian lineages infecting G. plebejus: Haemoproteus sp. GEPLE01 and GEPLE02, Haemoproteus sp. AMAVIR01, Haemoproteus sp. ZOCAP08, P. homopolare BAEBIC02 and P. cathemerium ZONCAP15. Lineages GEPLE01 and GEPLE02 are novel and are the 4th and 5th haemosporidian lineages reported for this species, after lineages Leucocytozoon ANIIGN02 and Haemoproteus CONCIN03 from Peru (McNew et al., Reference McNew, Barrow, Williamson, Galen, Skeen, DuBay, Gaffney, Johnson, Bautista, Ordoñez, Schmitt, Smiley, Valqui, Bates, Hackett and Witt2021), and PHRPLE01 from Chile (Doussang et al., Reference Doussang, Sallaberry-Pincheira, Cabanne, Lijtmaer, González-Acuña and Vianna2021). Our results suggest that GEPLE01 or GEPLE02 originated as a single, silent mutation of the other. Thus, differences in prevalence and mean parasitaemia are unrelated to their molecular identity for the cytb fragment analysed herein. According to the morphological data, these lineages could be associated with the unidentified Haemoproteus sp. 2 or Haemoproteus sp. 3 morphologies (Table 1). In-depth analyses must be performed to determine if they correspond to new species.
The most common lineage in this study was Haemoproteus sp. AMAVIR01. The only known host infected by Haemoproteus sp. AMAVIR01 is the hummingbird Amazilia viridicauda (GenBank: JQ988544) from Calca (Peru) at an elevation of 2953 m a.s.l. (McNew et al., Reference McNew, Barrow, Williamson, Galen, Skeen, DuBay, Gaffney, Johnson, Bautista, Ordoñez, Schmitt, Smiley, Valqui, Bates, Hackett and Witt2021; Dataset_S01). To date, no morphological identification has been provided for this lineage. According to our results, it is most likely associated with morphospecies H. coatneyi (Table 1). The capacity to infect Apodiformes and Passeriformes may indicate that this lineage is host-generalist, which is not common but has been reported for Haemoproteus (e.g. Moens et al., Reference Moens, Valkiūnas, Paca, Bonaccorso, Aguirre and Pérez-Tris2016). An alternate explanation is that since this lineage was detected in A. viridicauda only by sequencing (no microscopy was applied), this hummingbird might not be a competent host for Haemoproteus sp. AMAVIR01. A positive PCR diagnosis might result from sporozoite-stage infection and abortive development in the hosts (Valkiūnas et al., Reference Valkiūnas, Palinauskas, Ilgūnas, Bukauskaitė, Dimitrov, Bernotienė, Zehtindjiev, Ilieva and Iezhova2014). This possibility highlights the importance of combining microscopy and molecular diagnosis when studying avian haemosporidian parasites (Palinauskas et al., Reference Palinauskas, Žiegytė, Iezhova, Ilgūnas, Bernotienė and Valkiūnas2016).
The lineage Haemoproteus sp. ZOCAP08 and both Plasmodium lineages, ZONCAP15 and BAEBIC02, infect the rufous-collared sparrow, Zonotrichia capensis, in the study area (Cadena-Ortiz et al., Reference Cadena-Ortiz, Mantilla, Rivero de Aguilar, Flores, Bahamonde, Matta and Bonaccorso2019). Lineage ZOCAP08 infects at least 12 other avian species from North America to Argentina (Jones et al., Reference Jones, Cheviron and Carling2013; Reinoso-Pérez et al., Reference Reinoso-Pérez, Canales-Delgadillo, Chapa-Vargas and Riego-Ruiz2016; Carbó-Ramírez et al., Reference Carbó-Ramírez, Zuria, Schaefer and Santiago-Alarcon2017; Ham-Dueñas et al., Reference Ham-Dueñas, Chapa-Vargas, Stracey and Huber-Sannwald2017; Fecchio et al., Reference Fecchio, Bell, Pinheiro, Cueto, Gorosito, Lutz, Gaiotti, Paiva, França, Toledo-Lima, Tolentino, Pinho, Tkach, Fontana, Grande, Santillán, Caparroz, Roos, Bessa, Nogueira, Moura, Nolasco, Comiche, Kirchgatter, Guimarães, Dispoto, Marini, Weckstein, Batalha-Filho and Collins2019; Barrow et al., Reference Barrow, Bauernfeind, Cruz, Williamson, Wiley, Ford, Baumann, Brady, Chavez, Gadek, Galen, Johnson, Mapel, Marroquin-Flores, Martinez, McCullough, McLaughlin and Witt2021; McNew et al., Reference McNew, Barrow, Williamson, Galen, Skeen, DuBay, Gaffney, Johnson, Bautista, Ordoñez, Schmitt, Smiley, Valqui, Bates, Hackett and Witt2021) but has not been assigned a morphospecies in the MalAvi repository. To date, it has been attributed to H. coatneyi (González et al., Reference González, Lotta, García, Moncada and Matta2015) and Haemoproteus sp. (Carbó-Ramírez et al., Reference Carbó-Ramírez, Zuria, Schaefer and Santiago-Alarcon2017). Our study suggests an association with H. erythrogravidus and Haemoproteus sp. 1., but this result comes from a single positive sample that could have been infected by multiple morphospecies. Thus, more research is needed to support an association between Haemoproteus sp. ZOCAP08 and a specific morphology. Plasmodium cathemerium ZONCAP15 was found to be infecting 2 individuals with low parasitaemia. This lineage also showed low prevalence (2.26% by molecular diagnosis) in Z. capensis analysed for the same period and by the same methodology (Cadena-Ortiz et al., Reference Cadena-Ortiz, Mantilla, Rivero de Aguilar, Flores, Bahamonde, Matta and Bonaccorso2019). However, the only sample diagnosed by microscopy showed one of the highest parasitaemias found for that host (Cadena-Ortiz et al., Reference Cadena-Ortiz, Mantilla, Rivero de Aguilar, Flores, Bahamonde, Matta and Bonaccorso2019). Plasmodium cathemerium is a well-known generalist that infects many hosts in different avian taxa and is highly pathogenic (Vanstreels et al., Reference Vanstreels, da Silva-Filho, Kolesnikovas, Bhering, Ruoppolo, Epiphanio, Amaku, Junior, Braga and Catão-Dias2015), which could explain its low prevalence. Still, monitoring this parasite in other host species is needed to understand its prevalence and pathogenicity. Finally, lineage BAEBIC02 P. homopolare is widespread in the Americas, from Alaska to Peru, and has been found in at least 15 other avian species (Martinsen et al., Reference Martinsen, Waite and Schall2007; Galen and Witt, Reference Galen and Witt2014; Oakgrove et al., Reference Oakgrove, Harrigan, Loiseau, Guers, Seppi and Sehgal2014; Walther et al., Reference Walther, Valkiūnas, González, Matta, Ricklefs, Cornel and Sehgal2014; González et al., Reference González, Lotta, García, Moncada and Matta2015; Marzal et al., Reference Marzal, García-Longoria, Cardenas Callirgos and Sehgal2015). Finally, our recapture data, showing persistent haemosporidian infections, are valuable because of the lack of repetitive measures in field studies. However, a higher sample and serial recaptures would be necessary to analyse changes in host condition and parasitaemia during the course of the infection.
It is important to state that our study may be underestimating the diversity of haemosporidians in G. plebejus because we did not use the nested PCR approach of Hellgren et al. (Reference Hellgren, Waldenström and Bensch2004), which is the standard for current avian malaria studies. In their methodology, a fragment of haemosporidian cytb is amplified, and then a second PCR is applied on that fragment to amplify either Leucocytozoon, or Plasmodium and Haemoproteus. Here, we used only the second step for amplifying Plasmodium and Haemoproteus, as originally proposed by Bensch et al. (Reference Bensch, Stjernman, Hasselquist, Örjan, Hannson, Westerdahl and Pinheiro2000). Failing to use the nested PCR approach may have decreased the sensibility of the PCR assay in detecting several infections, especially low-intensity Plasmodium sp. infections (Waldenström et al., Reference Waldenström, Bensch, Hasselquist and Östman2004). This methodological approach may also be responsible for the small number of coinfections detected by PCR and at least some discrepancies between the molecular and microscopic diagnosis in detecting coinfections.
Morphological diagnosis of other haemoparasites
We found no evidence of Leucocytozoon infection. Based on the distribution of blackflies (Simuliidae), Lotta et al. (Reference Lotta, Pacheco, Escalante, González, Mantilla, Moncada, Adler and Matta2016) suggested that the transmission of this parasite is optimal above 2400 m a.s.l. but it may start as low as 2000 m a.s.l. This elevational limit seems to be related to the environmental conditions required by the life cycle of its vectors, blackflies (Simuliidae), in the Andes (Matta et al., Reference Matta, Lotta, Valkiūnas, González, Pacheco, Escalante, Moncada and Rodríguez-Fandiño2014). Our sample site lies between 2000 and 2500 m a.s.l., and 2 blackfly species occur in the area, 1 of them at a relatively high abundance (Subía-Solís, Reference Subía-Solís2013). Although we found no evidence of Leucocytozoon infection by microscopy, this parasite might persist at low parasitaemia and remain undetected. Thus, PCR amplification of the parasite DNA in avian hosts and blackflies of this community would be advisable in future studies.
We detected a microfilaria in 1 sample out of 65 samples diagnosed by microscopy (1.5% prevalence). The prevalence of microfilaria infection in Andean birds at the community level is relatively low, in the order of 0‒3% (2.3%, Bennett and Borrero, Reference Bennett and Borrero1976; 3%, Valkiūnas et al., Reference Valkiūnas, Salaman and Iezhova2003; 0%, Munro et al., Reference Munro, Martin, Moore and Bonier2009; 2.9%, Rodríguez et al., Reference Rodríguez, Moya and Matta2009). However, even if they show low prevalence, diagnosing and reporting microfilariae is important since they may interact with malarial infections, affecting the host's health condition (Clark et al., Reference Clark, Wells, Dimitrov and Clegg2016).
We also found 1 sample infected with Babesia (Apicomplexa: order Piroplasmida). Parasites in this genus are transmitted by ticks (order Ixodida) and are known to have zoonotic potential (Yabsley and Shock, Reference Yabsley and Shock2013; Shock et al., Reference Shock, Moncayo, Cohen, Mitchell, Williamson, Lopez, Garrison and Yabsley2014). Babesia species have been detected in several bird families, including Passeriformes and non-Passeriformes (Peirce, Reference Peirce2000; Yabsley et al., Reference Yabsley, Vanstreels, Shock, Purdee, Horne, Peirce and Parsons2017; Ebani and Mancianti, Reference Ebani and Mancianti2021). However, instances of infections in Neotropical birds have been poorly documented. There are reports of infection in Orinoco geese, Neochen jubata, in Brazil (Werther et al., Reference Werther, de Cássia Luzzi, Gonçalves, de Oliveira, Junior, Machado and André2017), and veery, Catharus fuscescens, in Canada (Scott et al., Reference Scott, Clark and Durden2019). To our knowledge, the only known Passerines captured in South America with a Babesia sp. infection are the Juan Fernández turdus, Turdus falcklandii, and Juan Fernández tit-tirant, Anairetes fernandezianus, from Juan Fernández archipelago, and T. falcklandii from mainland Chile (Martínez et al., Reference Martínez, Vásquez, Venegas and Merino2015). Woodworth-Lynas et al. (Reference Woodworth-Lynas, Caines and Bennett1989) reported Babesia in Brazil, but collectively, as part of ‘other’ parasitic genera; therefore, no specific information was provided. Thus, much research is needed to understand Babesia parasites in Passeriformes, especially in South America and the Neotropics.
Patterns of prevalence and parasitaemia
Molecular diagnosis revealed a high prevalence of haemosporidian infections in the population. This result can be related to the high abundance of the host in the study area, the 3rd most abundant in the community. The 1st and 2nd most abundant species are the Z. capensis and the common ground dove, Columbina passerina, which also show high haemosporidian prevalence and parasitaemia (Cadena-Ortiz et al., Reference Cadena-Ortiz, Mantilla, Rivero de Aguilar, Flores, Bahamonde, Matta and Bonaccorso2019; DB, HFC and EB, unpublished data). Considering the relative abundance of the host is important because prevalence usually increases alongside local host abundance (e.g. Ricklefs et al., Reference Ricklefs, Swanson, Fallon, Martínez-Abraín, Scheuerlein, Gray and Latta2005; Matthews et al., Reference Matthews, Ellis, Hanson, Roberts, Ricklefs and Collins2016). Still, a more comprehensive sampling of the avian community at this site is necessary to determine if this pattern holds.
Haemoproteus spp. were 4 times more prevalent than Plasmodium spp., which is consistent with previous studies (Bensch et al., Reference Bensch, Stjernman, Hasselquist, Örjan, Hannson, Westerdahl and Pinheiro2000; Clark et al., Reference Clark, Clegg and Lima2014), but the prevalence of Plasmodium could be particularly underestimated because of our choice of a non-nested PCR approach. However, even if we failed to detect several Plasmodium infections in the sample, the morphological identification also detected a low prevalence of Plasmodium. This result suggests that high-intensity infections by Plasmodium are rare or that affected individuals reduce their activity, lowering their capture probability.
Our results point to higher parasitaemia by Haemoproteus than Plasmodium, which is also consistent with previous studies (e.g. Fallon and Ricklefs, Reference Fallon and Ricklefs2008; Rodríguez-Hernández et al., Reference Rodríguez-Hernández, Álvarez-Mendizábal, Chapa-Vargas, Escobar, González-García and Santiago-Alarcon2021). Contrary to our expectations, we found no effect of infection, lineage or coinfection on body condition or polychromatophil count. Lack of differences in body condition have been observed between infected and non-infected individuals (Granthon and Williams, Reference Granthon and Williams2017), and, in the form of body weight, between single infections and coinfections with 2 different species and lineages of Plasmodium (Palinauskas et al., Reference Palinauskas, Žiegytė, Šengaut and Bernotienė2018, Reference Palinauskas, Žiegytė, Šengaut and Bernotienė2022). We expected that polychromatophil count would differ, at least between infected and non-infected individuals. However, since polychromasia increases with parasitaemia (Palinauskas et al., Reference Palinauskas, Žiegytė, Šengaut and Bernotienė2022), an individual with low parasitaemia may have similar levels of polychromasia as a non-infected one. Still, uncovering some of these relationships may be hampered by our methodological limitations in PCR diagnosis and relatively small sample size.
On the other hand, we found that parasitaemia was marginally higher in samples with coinfections than in samples with single infections. This trend coincides with the results of Palinauskas et al. (Reference Palinauskas, Žiegytė, Šengaut and Bernotienė2018) when simulating coinfections by 2 different species of Plasmodium. However, to understand the underlying processes that regulate parasitaemia of coinfections, more studies are needed on the interactions between different species and lineages of haemosporidians, and among them and the immune system of the host.
Predictors of prevalence and parasitaemia
The age of the host was the only predictor of infection status, with immature ones showing lower prevalence (63%) than adults (97%). These results are coherent with previous studies in Neotropical and temperate birds (e.g. Wood et al., Reference Wood, Cosgrove, Wilkin, Knowles, Day and Sheldon2007; Fecchio et al., Reference Fecchio, Lima, Silveira, Ribas, Caparroz and Marini2015; Cadena-Ortiz et al., Reference Cadena-Ortiz, Mantilla, Rivero de Aguilar, Flores, Bahamonde, Matta and Bonaccorso2019). Prevalence may increase with age as survivors acquire immunity to the parasite (see Atkinson et al., Reference Atkinson, Dusek and Lease2001). However, the relationship between age and prevalence must be context-dependent, in terms of both the environment and natural history of the host, since other studies have found that prevalence increases with age (e.g. van Oers et al., Reference van Oers, Richardson, Sæther and Komdeur2010; Hammers et al., Reference Hammers, Komdeur, Kingma, Hutchings, Fairfield, Gilroy and Richardson2016).
Parasitaemia, on the other hand, was not predicted by any of the variables included in the GLMs. Negative results for all but 1 predictor of prevalence and all predictors of parasitaemia might result from our relatively small sampling effort because of 2 main reasons. First, although our sampling size might be adequate for other species, it might not be for species with a high prevalence of infection. When non-infected individuals are rare, parameter value estimation for these individuals is challenging. Second, parasitaemia variance is naturally high, which may also complicate parameter estimation with relatively reduced sample sizes. Still, some additional factors should be considered to better understand this system. On the other hand, we found no effect of precipitation on prevalence and parasitaemia, which was surprising, considering that there was variation in total precipitation in the month before the capture day, from 16.7 to 91.8 mm, more than a 5-fold difference. Cadena-Ortiz et al. (Reference Cadena-Ortiz, Mantilla, Rivero de Aguilar, Flores, Bahamonde, Matta and Bonaccorso2019) found an effect of precipitation on haemosporidian prevalence for Z. capensis in the same area, using the same sampling and parasite-screening methodology. This difference might be explained by host ecology, physiology and differences among infecting parasites, or, again, a higher sampling size (i.e. 177 individuals).
We also found no effect of host sex on prevalence or parasitaemia, which might result from an interplay of several factors. Males of G. plebejus may maintain secondary sexual characters at an immunological cost (Hamilton and Zuk, Reference Hamilton and Zuk1982; Zuk, Reference Zuk1990), although this species' dimorphism is moderate (see plates in Jaramillo, Reference Jaramillo2021). On the other hand, females may pay a similar or higher cost by exposing themselves to vector bites during incubation. Still, more information on the reproductive behaviour of males and females of this species is necessary to posit and test more informed hypotheses.
Finally, other factors not measured herein might predict prevalence and parasitaemia better. First, given the seasonality of the study area, a year-long analysis is necessary to determine more adequately the relationship between malarial infection and environmental factors. Second, other variables such as the abundance of and distance to water sources, vegetation structure and remote-sensing derived data (e.g. normalized difference vegetation index) may help documenting the heterogeneity among sites (e.g. Hernández-Lara et al., Reference Hernández-Lara, González-García and Santiago-Alarcon2017; Ferraguti et al., Reference Ferraguti, Martínez-de la Puente, Bensch, Roiz, Ruiz, Viana, Soriguer and Figuerola2018). Also, additional measures of health condition (e.g. heterophil/lymphocyte ratio, haematocrit level), and individuals' reproductive status may generate more robust predictions of both prevalence and parasitaemia.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0031182022001603.
Data availability
Unique DNA sequences for haemosporidian parasites were uploaded to GenBank under accession nos. ON938203 and ON938204 (https://www.ncbi.nlm.nih.gov/genbank/). Raw data are available in Supplementary File 1.
Acknowledgements
We are grateful to Nicolás Peñafiel for processing a portion of the molecular samples and supervising molecular work, and to Ingrid Lotta for her technical assistance and help in creating the tables. Ralph Vanstreels provided valuable literature and insights on Babesia records, and Tomas Roque helped editing the English language. Consejo Provincial de Pichincha provided access to the study area and Ministerio de Ambiente del Ecuador authorized this study through permiso de investigación no. 20-2012-IC-FAU-DPAP-MA and Contrato de Acceso a Recursos Genéticos MAE-DNB-CM-2018-0105 granted to Universidad Tecnológica Indoamérica. We acknowledge INAMHI for providing climate data for statistical analyses.
Author's contributions
X. C., H. C.-O., N. E. M., I. A. and E. B. conceived the study; H. C.-O. obtained the samples in the field; E. B. supervised molecular diagnosis and X. C. performed phylogenetic analyses; I. A. and D. B.-V. generated data on parasitaemia; D. B.-V. generated molecular data; N. E. M. and A. D. G. performed morphological determination; X. C. performed all statistical analyses; X. C., E. B. and N. E. M. wrote the first draft of the manuscript. All authors contributed to previous versions and read and approved the final version of the manuscript.
Financial support
This study was supported by Universidad Tecnológica Indoamérica (Convocatoria a Proyectos 2012) and Universidad San Francisco de Quito (project HUBi 12434).
Conflict of interest
The authors declare no conflict of interests.
Ethical standards
During the field phase of this study, the handling of birds maintained the highest ethical and legal standards.