Introduction of main epigenetic processes in psychiatry research
Both genetic and environmental factors are implicated in the aetiology of psychiatric disorders, however, the key causal mechanisms for guiding effective prevention and treatment remain poorly understood (Van Os, Rutten, & Poulton, Reference Van Os, Rutten and Poulton2008). Genetic association studies (Ripke et al., Reference Ripke, Neale, Corvin, Walters, Farh, Holmans and Huang2014) as well as epidemiological studies addressing the impact of the environment (van Os, Kenis, & Rutten, Reference van Os, Kenis and Rutten2010) on disease burden, have not yet explained the non-complete genetic correlation between monozygotic twins in conditions such as schizophrenia (SCZ) (41–65%), Bipolar Disorder (BD) (~60%) (Craddock, O'Donovan, & Owen, Reference Craddock, O'Donovan and Owen2005) or Major Depression (MDD) (~40%) (Ripke et al., Reference Ripke, Wray, Lewis, Hamilton, Weissman, Breen and Cichon2013).
In the past decade, growing evidence has shown a link between epigenetic processes, and a range of mental health disorders (Binder, Reference Binder2017). Epigenetic modifications refer to functional changes in DNA structural packaging or associated proteins without structural alteration of the DNA sequence itself (Jaenisch & Bird, Reference Jaenisch and Bird2003). This biological mechanism has important implications on how genes are expressed and how the chromatin is packaged, thus modifying subsequent protein translation within regionally specific parts of the central nervous system (Binder, Reference Binder2017). The most studied epigenetic process in humans is DNA methylation (DNAm) (Table 1 for definitions of key terms). Indeed, recent parallel evidence suggests that differential DNAm profiles are associated with exposure to childhood adversity (CA) as well as cannabis use (CU) (Kandaswamy et al., Reference Kandaswamy, Hannon, Arseneault, Mansell, Sugden, Williams and Wong2020; Markunas et al., Reference Markunas, Hancock, Xu, Quach, Fang, Sandler and Taylor2020; Nöthling, Malan-Müller, Abrahams, Hemmings, & Seedat, Reference Nöthling, Malan-Müller, Abrahams, Hemmings and Seedat2020). This suggests that epigenetic factors may account for some of the non-explained variance in genetics studies and possibly mediate the interactions between genotype and known environmental risk factors in influencing the onset of complex diseases (Relton & Smith, Reference Relton and Smith2010).
Table 1. A glossary of key epigenetic terms and biological function of genes involved in pathways discussed in this review
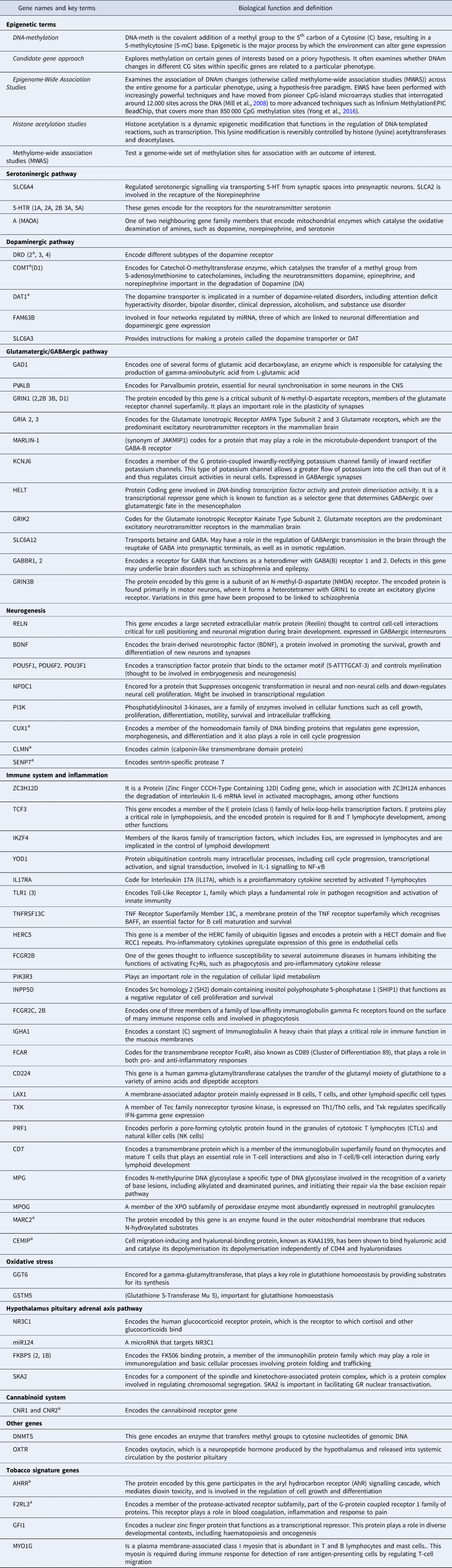
a genes related to cannabis use.
Initially, epigenetic research in psychiatry used a candidate gene approach, and progressively, research moved to Epigenome Wide Association studies (EWAS) (Table 1). While both designs have their advantages and limitations, the breadth of coverage of EWAS offers a more informative insight on biological pathways. This is based on the rational that chromatin conformation and transcriptional regulation are influenced by a set of methylated or unmethylated cytosines across a region, rather than specific CpG sites in isolation (Mill et al., Reference Mill, Tang, Kaminsky, Khare, Yazdanpanah, Bouchard and Petronis2008)
Different biological pathways have been implicated in the aetio-pathogenesis across multiple mental disorders. Some of these are pathways related to neurotransmission such as serotonin (Provenzi, Giorda, Beri, & Montirosso, Reference Provenzi, Giorda, Beri and Montirosso2016), dopamine or GABA/glutamatergic processes (McCutcheon, Krystal, & Howes, Reference McCutcheon, Krystal and Howes2020); while others pathways involve inflammation (Cullen et al., Reference Cullen, Holmes, Pollak, Blackman, Joyce, Kempton and Mondelli2019), oxidative stress (Steullet et al., Reference Steullet, Cabungcal, Monin, Dwir, O'Donnell, Cuenod and Do2016), synaptic plasticity and neurogenesis (Claudino, Gonçalves, Schuch, Martins, & Rocha, Reference Claudino, Gonçalves, Schuch, Martins and Rocha2020), or the stress response system (Hypothalamic Pituitary adrenal Axis – HPA) (Wesarg, Van Den Akker, Oei, Hoeve, & Wiers, Reference Wesarg, Van Den Akker, Oei, Hoeve and Wiers2020). It is important to take into account that some of these processes participate in disease pathogenesis in a parallel manner, such as via the redox system and through the glutamatergic/GABAergic imbalance (Hardingham & Do, Reference Hardingham and Do2016); or the immune system and the stress response (Pariante, Reference Pariante2017). Although these processes are often explored within discrete categorical clinical conditions, they often overlap transdiagnostically. For instance, alterations in serotonin pathways are linked to both depression and psychosis phenotypes (Selvaraj, Arnone, Cappai, & Howes, Reference Selvaraj, Arnone, Cappai and Howes2014).
In this review, we set to appraise firstly, the evidence of DNAm modifications both from candidate genes and EWAS studies, associated either specifically or transdiagnostically with psychiatric conditions, and secondly, if these DNAm modifications map onto biological pathways. Thirdly, we will explore if the existing findings from studies on DNAm changes associated with CA and CU, two of the environmental exposures most consistently associated with psychiatric disorders (Lindert et al., Reference Lindert, von Ehrenstein, Grashow, Gal, Braehler and Weisskopf2014; Mandelli, Petrelli, & Serretti, Reference Mandelli, Petrelli and Serretti2015; Marconi, Di Forti, Lewis, Murray, & Vassos, Reference Marconi, Di Forti, Lewis, Murray and Vassos2016; Sideli, Quigley, La Cascia, & Murray, Reference Sideli, Quigley, La Cascia and Murray2020a; Varese et al., Reference Varese, Smeets, Drukker, Lieverse, Lataster, Viechtbauer and Bentall2012), point at the same biological pathways therefore contributing to the understanding of how these environmental exposures increase transdiagnostic and specific psychiatric liability. Details on methodological considerations can be found in Online Supplementary Material (SM).
Evidence of epigenetic processes in major transdiagnostic pathways
In this section we will review the evidence, predominantly from case–control studies pointing at an association between DNAm changes in the Serotoninergic, Dopaminergic pathways, Excitatory inhibitory balance (including the Glutamatergic and GABAergic dysfunction), Synaptic plasticity and Neurogenesis; the Immune system, Inflammation and Oxidative stress and the major mental conditions (focusing on Eating Disorders (ED): anorexia nervosa (AN) and bulimia nervosa (BN), Autism Spectrum Disorder (ASD), BD and Psychotic Disorder, Depression, Post Traumatic Stress Disorder (PTSD) and Anxiety Disorders). Summary of findings is illustrated in Fig. 1; findings on HPA-axis and its association to environmental risk factors are presented in Section ‘The epigenetic signature of childhood adversity and cannabis use’ and Fig. 2 and Online Supplementary Table S1 (SM) summarises the characteristics of the articles mentioned in that section. Table 2 summarised the key elements of studies finding evidence of a link between DNAm on genes involved in each biological pathway across all disorders. Screening

Fig. 1. Summary of the evidence on potential pathways linking childhood trauma and cannabis use with psychiatric conditions through DNAm changes.
Note: This figure summarises the evidence presented in this review, highlighting the idea that some biological pathways linking environmental risk factors with mental health disorders via epigenetic changed in the form of DNAm are transdiagnostics (e.g immune system/inflammation) while others seem to be more specific (e.g dopaminergic system). (1) The environmental risk factors row and epigenetic modifications row suggest links between childhood adversity (CA), and Cannabis use (CU) and DNAm changes mapping to biological pathways which are also functionally related (Serotoninergic, Dopaminergic pathways, Glutamatergic & GABAergic pathway, Neurogenesis, Immune system & Inflammation and Oxidative stress). (2)The epigenetic modifications row and mental health disorders row illustrate the evidence, from case–control studies, of an association between DNAm changes in these pathways and the major mental health conditions (Eating Disorders (anorexia nervosa and bulimia nervosa) Post-traumatic stress disorder, Anxiety Disorders, Psychotic Disorder, Bipolar disorders, Depression and Autism Spectrum Disorders). (3) The arrows connecting the three rows show the potential mediating role of DNAm changes linking CA and CU and risk to develop mental health conditions. The thickness of the lines shows the robustness of the evidence reported in the literature review. The items “genotype: and “other risk factors” are added to highlight the influence of genetic factors and environmental confounders in DNAm studies. The dotted line connecting eating disorders with the pathways indicate that literature was limited and mixed not allowing to draw clear links with the pathways.

Fig. 2. Summary of the evidence linking childhood adversity and DNAm changes on the Hipotalamic Pituitary Adrenal Axis in various conditions as well as with some clinical measures.
Note: This Figure illustrates the evidence from candidate gene studies linking childhood adversity (CA) with DNAm in CpG sites located in NR3C1, FKBP5, SKA2 and CA, with various conditions and various clinical outcomes. In the gene and DNAm columns, CA + (with an arrow pointing up) reflects the presence of a positive association between the DNAm in probes located in those genes and CA; CA- (with an arrow pointing down) reflects a negative association. The disorder column shows in which mental health condition that association has been found. Lastly, the clinical outcomes column shows the presence of evidence linking DNAm, with a particular clinical phenotype; CA + indicated that the association between DNAm and the clinical outcome was related to CA.
Table 2. Summary of the direction of the associations between DNAm, mental health disorders and clinical or biological outcomes presented in this review

*extensive reviews cover the role of BDNF Methylation in depression (Hing et al., Reference Hing, Sathyaputri and Potash2018), Schizophrenia (Di Carlo et al., Reference Di Carlo, Punzi and Ursini2019), and eating disorders (Thaler and Steiger, Reference Thaler and Steiger2017), therefore studies mentioned here are just examples of the literature in this particular domain. When various genes are reported in the same pathway and the same study, but no specific information on clinical/biological outcome or specific direction if the association is provided, these genes have been put in the same row (e.g Asberg et al., and Liu et al.,). When an arrow is next to the author's name it reflects the direction of the DNAm of the particular gene in in relation to the condition ↑ : increased ↓: decreased DNAm. When in column 1 there is no arrow is because information could not be obtained or was not clear, and the presence of that gene indicates the association of DNAm in that gene with the respective condition (differently methylated). When a three step sequence separated by an arrow is presented, this refers to mediation analyses (e.g peng et al.,: CA → ↑SLC6A4 → depressive symptoms: DNAm of SLC6A mediates the effect of CA on depressive symptoms). CA: childhood adversity; CU: cannabis use. Definition of each gene is presented in Table 1. DLPFC: Dorsolateral prefrontal cortex. ASD: autism spectrum disorder; SCZ: schizophrenia. PTSD: post-traumatic stress disorder; Borderline PD: Borderline personality disorder; MDD: major depression disorder; BD: bipolar disorder.
The serotoninergic pathway
There are preclinical and human studies pointing at an implication of the serotonin (5HT) system dysfunction in a broad range of psychiatric diseases (Kaye, Fudge, & Paulus, Reference Kaye, Fudge and Paulus2009). The strongest evidence is at the level of the serotonin transported genes (mainly SLC6A4) with candidate genes studies suggesting an increased in methylation in depression (Kang et al., Reference Kang, Kim, Stewart, Kim, Bae, Kim and Yoon2013; Philibert et al., Reference Philibert, Sandhu, Hollenbeck, Gunter, Adams and Madan2008; Zhao, Goldberg, Bremner, & Vaccarino, Reference Zhao, Goldberg, Bremner and Vaccarino2013), BD (Sugawara et al., Reference Sugawara, Iwamoto, Bundo, Ueda, Miyauchi, Komori and Okazaki2011) and reporting a positive association with symptoms severity (Olsson et al., Reference Olsson, Foley, Parkinson-Bates, Byrnes, McKenzie, Patton and Saffery2010), comorbid depression in those with panic disorder (Schiele et al., Reference Schiele, Kollert, Lesch, Arolt, Zwanzger, Deckert and Domschke2019), and improvement from baseline to follow-up (Perez-Cornago, Mansego, Zulet, & Martinez, Reference Perez-Cornago, Mansego, Zulet and Martinez2014). It has been suggested that an increased DNAm of SLC6A4 could repress gene expression, leading to decreased serotonin uptake and lower activity, which ultimately would lead to the manifestation of depressive symptoms (Chen, Meng, Pei, Zheng, & Leng, Reference Chen, Meng, Pei, Zheng and Leng2017)
A pattern of hypermethylation has also been found in samples of SCZ (Abdolmaleky et al., Reference Abdolmaleky, Nohesara, Ghadirivasfi, Lambert, Ahmadkhaniha, Ozturk and Thiagalingam2014). although with mixed evidence(Alelú-Paz et al., Reference Alelú-Paz, González-Corpas, Ashour, Escanilla, Monje, Guerrero Marquez and Ropero2015). Candidate gene studies in SCZ and BD across various tissues (Abdolmaleky et al., Reference Abdolmaleky, Yaqubi, Papageorgis, Lambert, Ozturk, Sivaraman and Thiagalingam2011; Carrard, Salzmann, Malafosse, & Karege, Reference Carrard, Salzmann, Malafosse and Karege2011; Cheah, Lawford, Young, Morris, & Voisey, Reference Cheah, Lawford, Young, Morris and Voisey2017) show elevated DNAm of the 5-HTR1A and 5-HTR2A genes respectively. Further, EWAS studies have identified differential DNAm in HTR2A (Numata, Ye, Herman, & Lipska, Reference Numata, Ye, Herman and Lipska2014), HTR5A and HTR1E (Nishioka et al., Reference Nishioka, Bundo, Koike, Takizawa, Kakiuchi, Araki and Iwamoto2013; Pidsley et al., Reference Pidsley, Viana, Hannon, Spiers, Troakes, Al-Saraj and Bray2014) genes in those with psychosis.
Evidence on ED so far has not found an association with SLC6A4 DNAm and AN (Boehm et al., Reference Boehm, Walton, Alexander, Batury, Seidel, Geisler and Ehrlich2019; Pjetri et al., Reference Pjetri, Dempster, Collier, Treasure, Kas, Mill and Schmidt2013; Steiger et al., Reference Steiger, Booij, Kahan, McGregor, Thaler, Fletcher and Szyf2019).
In ASD, preliminary evidence indicated higher HTR2A promoter DNAm in leucocytes of those carrying the high-risk genotype in the HTR2A (Hranilovic, Blazevic, Stefulj, & Zill, Reference Hranilovic, Blazevic, Stefulj and Zill2016).
Another well-explored gene of interest in the serotoninergic pathway is MAO-A (Shih & Thompson, Reference Shih and Thompson1999) which is involved in monoamine degradation and it has established linked with depression (Meyer et al., Reference Meyer, Ginovart, Boovariwala, Sagrati, Hussey, Garcia and Houle2006). While studies in depression have found inconsistent DNAm changes (Domschke et al., Reference Domschke, Tidow, Schwarte, Ziegler, Lesch, Deckert and Baune2015; Melas & Forsell, Reference Melas and Forsell2015; Melas et al., Reference Melas, Wei, Wong, Sjöholm, Åberg, Mill and Lavebratt2013); in candidate gene studies in anxiety disorders, the evidence points at a pattern of hypomethylation (Ziegler et al., Reference Ziegler, Richter, Mahr, Gajewska, Schiele, Gehrmann and Helbig-Lang2016) as shown in acrophobia (Schiele et al., Reference Schiele, Ziegler, Kollert, Katzorke, Schartner, Busch and Deckert2018) and obsessive compulsive disorder (OCD) (Domschke et al., Reference Domschke, Tidow, Kuithan, Schwarte, Klauke, Ambrée and Kersting2012). Moreover, increased MAO-A DNAm has been suggested as a potential useful marker of better response to psychotherapy in anxiety disorders (Schiele et al., Reference Schiele, Thiel, Deckert, Zaudig, Berberich and Domschke2020; Ziegler et al., Reference Ziegler, Richter, Mahr, Gajewska, Schiele, Gehrmann and Helbig-Lang2016).
Overall, we find a transdiagnostic link between DNAm changes in genes involved in the serotoninergic pathway, with limited evidence in ED (findings on PTSD discussed in Section ‘The epigenetic signature of childhood adversity and cannabis use’ and described in Table 2).
The dopaminergic pathway
It is widely accepted that dopaminergic dysregulation stands as one of the most supported hypotheses for the pathogenesis of SCZ and psychosis as a whole (Jauhar et al., Reference Jauhar, McCutcheon, Borgan, Veronese, Nour, Pepper and Howes2018; McCutcheon et al., Reference McCutcheon, Krystal and Howes2020). Studies examining DNAm in the blood of patients with SCZ as compared with controls have showed both higher and lower DNAm levels in different DA receptor's genes; with decreased DNAm in DRD3 (Dai et al., Reference Dai, Cheng, Zhou, Lv, Zhuang, Zheng and Duan2014) and DRD4 (Cheng et al., Reference Cheng, Wang, Zhou, Wang, Li, Zhuang and Dai2014); and in other dopamine receptors (Kordi-Tamandani, Sahranavard, & Torkamanzehi, Reference Kordi-Tamandani, Sahranavard and Torkamanzehi2013b; Yoshino et al., Reference Yoshino, Kawabe, Mori, Mori, Yamazaki, Numata and Ohmori2016).
Hypomethylation of the COMT membrane-bound isoform, has been identified in samples of people with SCZ across tissues (Abdolmaleky et al., Reference Abdolmaleky, Cheng, Faraone, Wilcox, Glatt, Gao and Carnevale2006; Nohesara et al., Reference Nohesara, Ghadirivasfi, Mostafavi, Eskandari, Ahmadkhaniha, Thiagalingam and Abdolmaleky2011; Nour El Huda et al., Reference Nour El Huda, Norsidah, Nabil Fikri, Hanisah, Kartini and Norlelawati2018; Walton et al., Reference Walton, Liu, Hass, White, Scholz, Roessner and Ehrlich2014), while the soluble isoform (S-COMT) has been reported to be hypermethylated (Melas et al., Reference Melas, Rogdaki, Ösby, Schalling, Lavebratt and Ekström2012; Murphy, O'Reilly, & Singh, Reference Murphy, O'Reilly and Singh2005). EWAS studies comparing SCZ patients with controls have found hypomethylation of SLC6A3, a dopamine transporter (Nishioka et al., Reference Nishioka, Bundo, Koike, Takizawa, Kakiuchi, Araki and Iwamoto2013), COMTD1 and FAM63B, a gene linked to dopaminergic gene expression (Aberg et al., Reference Aberg, McClay, Nerella, Clark, Kumar, Chen and Gao2014).
In ED, findings of DNAm changes affecting dopaminergic genes DAT and DRD2 are mixed (Frieling et al., Reference Frieling, Romer, Scholz, Mittelbach, Wilhelm, De Zwaan and Bleich2010; Pjetri et al., Reference Pjetri, Dempster, Collier, Treasure, Kas, Mill and Schmidt2013). It has been suggested that DNAm variation in the dopamine pathway in ED may be related to comorbid Borderline Personality Disorder (Borderline PD) (Groleau et al., Reference Groleau, Joober, Israel, Zeramdini, DeGuzman and Steiger2014) and exposure to CA (Section ‘The epigenetic signature of childhood adversity and cannabis use’ and Online Supplementary Table S1 (SM)).
None of the EWAS studies conducted in ASD has found evidence supporting an association with DNAm changes involved in the Dopaminergic pathway.
Overall, recent findings support a link between DNAm changes in genes involved in the dopaminergic pathway related to neurodevelopmental disorders such as SCZ, with limited evidence suggesting a link with other conditions.
Glutamatergic/GABAergic Pathway and excitatory/inhibitory balance
Alterations in glutamatergic and GABAergic pathways, which can lead to either excitatory/inhibitory imbalance, have been reported to play a role in the etiopathogenesis of psychotic disorders (McCutcheon et al., Reference McCutcheon, Krystal and Howes2020) and ASD (Marotta et al., Reference Marotta, Risoleo, Messina, Parisi, Carotenuto, Vetri and Roccella2020). Furthermore, N-Methyl-D-aspartic acid or N-Methyl-D-aspartate (NMDAR) hypofunction as well as a decrease in the parvalbumin-expressing fast-spiking interneurons (PVI), both processes being essential for the excitatory/inhibitory balance, have been widely shown to be involved in psychotic disorders (Thuné, Recasens, & Uhlhaas, Reference Thuné, Recasens and Uhlhaas2016).
In SCZ and psychosis, there is evidence from candidate genes studies across tissues supporting DNAm differences between cases and controls in genes such as the Parvalbumin (PVALB) gene(Fachim, Srisawat, Dalton, & Reynolds, Reference Fachim, Srisawat, Dalton and Reynolds2018), GMR2 and GMR5 of the glutamatergic receptors (Kordi-Tamandani, Dahmardeh, & Torkamanzehi, Reference Kordi-Tamandani, Dahmardeh and Torkamanzehi2013a); various CpG sites in the β2 subunit of the GABAa receptor gene (GABRB2) (Pun et al., Reference Pun, Zhao, Lo, Ng, Tsang, Nimgaonkar and Xue2011; Zong et al., Reference Zong, Zhou, Hou, Zhang, Jiang, Zhang and Deng2017), and in GRIN2B, involved in the function of NMDAR (Fachim et al., Reference Fachim, Loureiro, Corsi-Zuelli, Shuhama, Louzada-Junior, Menezes and Reynolds2019). A dysregulation of multiple DNAm positions in the regulatory network of GAD1, was identified in patients with SCZ and BD compared to controls (Ruzicka, Subburaju, & Benes, Reference Ruzicka, Subburaju and Benes2015).
In terms of EWAS Mill and colleagues (Mill et al., Reference Mill, Tang, Kaminsky, Khare, Yazdanpanah, Bouchard and Petronis2008) performed the first EWAS in post-mortem brains of SCZ and BD subjects compared to controls, and found DNAm changes associated with SCZ and BD at loci involved in glutamatergic (GRIA 2, GRIND3B) and GABAergic (MARLIN-1, KCNJ6, HELT) neurotransmission, supporting previous candidate genes results. Findings related to GRIA family genes have been replicated in latter EWAS studies (Aberg et al., Reference Aberg, McClay, Nerella, Xie, Clark, Hudson and Hultman2012; Numata et al., Reference Numata, Ye, Herman and Lipska2014), and other EWAS studies have confirmed DNAm changes in genes involving GABAergic neurotransmission (SLC6A12 and GABBR1) (Hannon et al., Reference Hannon, Dempster, Mansell, Burrage, Bass, Bohlken and Mill2021).
In ASD, an EWAS study on histone acetylation in participants with the disorder compared to controls found an enrichment of hyperacetylated sites in genes involved in GABA receptor activity (Ramaswami et al., Reference Ramaswami, Won, Gandal, Haney, Wang, Wong and Geschwind2020), although this has not been previously found on DNAm (Wong et al., Reference Wong, Smith, Hannon, Ramaswami, Parikshak, Assary and Sun2018).
Lastly, a Depression EWAS (Nagy et al., Reference Nagy, Suderman, Yang, Szyf, Mechawar, Ernst and Turecki2015) of post-mortem brains of suicide victims and controls found 115 differentially methylated regions (DMRs), which included regions related to GRIK2.
Overall, there is evidence linking DNAm changes on genes involved in the glutamatergic pathway mainly with psychosis, with some evidence suggesting a link with ASD and MDD
Synaptic plasticity and neurogenesis
Synaptic plasticity anomalies are associated with psychiatric conditions and may account for various symptoms, such as cognitive deficits (Claudino et al., Reference Claudino, Gonçalves, Schuch, Martins and Rocha2020; Di Carlo, Punzi, & Ursini, Reference Di Carlo, Punzi and Ursini2019; Lin & Huang, Reference Lin and Huang2020).
RELN is a good studies gene that has been linked to SCZ (Costa et al., Reference Costa, Chen, Davis, Dong, Noh, Tremolizzo and Guidotti2002). An aberrant DNAm status in RELN has been found in SCZ and BD patient as compared with controls (Fikri et al., Reference Fikri, Norlelawati, El-Huda, Hanisah, Kartini, Norsidah and Zamzila2017; Tamura, Kunugi, Ohashi, & Hohjoh, Reference Tamura, Kunugi, Ohashi and Hohjoh2007). Interestingly, peripheral blood hypomethylation in the RELN promoter was associated with poor cognitive functioning (Alfimova, Kondratiev, Golov, & Golimbet, Reference Alfimova, Kondratiev, Golov and Golimbet2018).
In ASD, an EWAS study in post-mortem brain found dysregulation in the pathway of phosphatidylinositol 3-kinase (PI3K) activity (Wong et al., Reference Wong, Smith, Hannon, Ramaswami, Parikshak, Assary and Mill2019), an enzyme that is involved in cellular growth, proliferation and differentiation, and which has been previously been associated with SCZ (Law et al., Reference Law, Wang, Sei, O'Donnell, Piantadosi, Papaleo and Vakkalanka2012).
Brain-derived Neurotrophic factor (BDNF) is essential for neurogenesis and extensively studied as a biomarker in psychiatry (Lin & Huang, Reference Lin and Huang2020) There is extensive evidence of a difference in DNAm in BDNF, as well as EWAS studies showing enrichment for the neurogenesis pathway in SCZ (Di Carlo et al., Reference Di Carlo, Punzi and Ursini2019; Ursini et al., Reference Ursini, Cavalleri, Fazio, Angrisano, Iacovelli, Porcelli and Gelao2016), BD (Dell et al., Reference Dell, Palazzo, Benatti, Camuri, Galimberti, Fenoglio and Altamura2014; Duffy et al., Reference Duffy, Goodday, Keown-Stoneman, Scotti, Maitra, Nagy and Turecki2019), PTSD (Kim et al., Reference Kim, Kim, Chung, Choi, Kim and Kang2017; Uddin et al., Reference Uddin, Aiello, Wildman, Koenen, Pawelec, de Los Santos and Galea2010), Depression (Hing, Sathyaputri, & Potash, Reference Hing, Sathyaputri and Potash2018; Kang et al., Reference Kang, Kim, Bae, Kim, Shin, Kim and Yoon2015), Borderline PD (Arranz et al., Reference Arranz, Gallego-Fabrega, Martín-Blanco, Soler, Elices, Dominguez-Clavé and Pascual2021; Thomas et al., Reference Thomas, Knoblich, Wallisch, Glowacz, Becker-Sadzio, Gundel and Nieratschker2018), Anxiety Disorders (D'Addario et al., Reference D'Addario, Bellia, Benatti, Grancini, Vismara, Pucci and Fenoglio2019), ED (Thaler et al., Reference Thaler, Gauvin, Joober, Groleau, Guzman and Ambalavanan2014), ASD (Ramaswami et al., Reference Ramaswami, Won, Gandal, Haney, Wang, Wong and Geschwind2020) thus making a well-replicated epigenetic transdiagnostic finding in psychiatry.
As a whole, transdiagnostic evidence suggests an involvement of DNAm changes in neurogenesis an neural plasticity.
Immune system and inflammation
Abundant evidence supports the role of neuroinflammation and altered immune processes in the aetiopathogeneses of various mental conditions (Mazza, Lucchi, Rossetti, & Clerici, Reference Mazza, Lucchi, Rossetti and Clerici2020; Pariante, Reference Pariante2017).
An EWAS by Montano et al. (Montano et al. Reference Montano, Taub, Jaffe, Briem, Feinberg, Trygvadottir and Feinberg2016) found differences in DNAm in genes involved in T-cell development in the blood of SCZ patients (ZC3H12D, TCF3, and IKZF4); other EWAS have also reported an enrichment in the immune system pathway by differently methylated genes (FR2B, PIK3R3, INPP5D, FCGR2C, IGHA1, FCAR; CD224, LAX1, TXK, PRF1, CD7, MPG, and MPOG) (Aberg et al., Reference Aberg, McClay, Nerella, Clark, Kumar, Chen and Gao2014; Hannon et al., Reference Hannon, Dempster, Viana, Burrage, Smith, Macdonald and Mill2016; Liu et al., Reference Liu, Chen, Ehrlich, Walton, White, Perrone-Bizzozero and Calhoun2014).
In depression, a discordant monozygotic twin study based on peripheral blood, found 39 DMRs associated to a lifetime history of MDD, which were significantly enriched in biological pathways associated to cytokine secretion (Zhu et al., Reference Zhu, Strachan, Fowler, Bacus, Roy-Byrne and Zhao2020). Another EWAS on post-mortem brain of people with late-MDD (Hüls et al., Reference Hüls, Robins, Conneely, De Jager, Bennett, Epstein and Wingo2020), found altered DNAm in the YOD1 locus, which is dysregulated in depression (Howren, Lamkin, & Suls, Reference Howren, Lamkin and Suls2009) and its implicated in the regulation of inflammatory processes (Schimmack et al., Reference Schimmack, Schorpp, Kutzner, Gehring, Brenke, Hadian and Krappmann2017).
Two EWAS studies in PTSD found differences in DNAm across genes part of biological pathways involved with inflammation and immune response (Kuan et al., Reference Kuan, Waszczuk, Kotov, Marsit, Guffanti, Gonzalez and Luft2017; Uddin et al., Reference Uddin, Aiello, Wildman, Koenen, Pawelec, de Los Santos and Galea2010).
In ASD, various EWAS studies have pointed at dysregulation of pathways related to immune response (Ramaswami et al., Reference Ramaswami, Won, Gandal, Haney, Wang, Wong and Geschwind2020; Wong et al., Reference Wong, Smith, Hannon, Ramaswami, Parikshak, Assary and Mill2019), and in a genome-wide microRNA (miRNA) expression profiling study (Wu, Parikshak, Belgard, & Geschwind, Reference Wu, Parikshak, Belgard and Geschwind2016).
Furthermore, an EWAS study from patients suffering from Panic Disorder found enrichment in genes involved in the regulation of lymphocytes (Shimada-Sugimoto et al., Reference Shimada-Sugimoto, Otowa, Miyagawa, Umekage, Kawamura, Bundo and Kaiya2017).
We can conclude that there is transdiagnostic, rather than specific, a link between DNAm changes in the immune system and inflammation and neural plasticity, although evidence is more robust in SCZ.
Oxidative stress
There is converging evidence pointing at a role of redox dysregulation as a possible mechanism involved in the aetiopathogenesis of both ASD (Bjørklund et al., Reference Bjørklund, Meguid, El-Bana, Tinkov, Saad, Dadar and Kizek2020) and psychosis (Perkins, Jeffries, & Do, Reference Perkins, Jeffries and Do2020). Oxidative stress has been shown to play a role in epigenetic modifications, enhancing inflammatory gene transcription (Rahman, Marwick, & Kirkham, Reference Rahman, Marwick and Kirkham2004). Oxidation of 5mC to the 5-hydroxymethylcytosine (5hmC) is considered a key step in the reversibility of DNA methylation, which can have important therapeutic implications. Moreover, glutathione, the major antioxidant in the brain, is involved in the methionine cycle, and depletion of glutathione can be detrimental for the DNAm process (García-Giménez, Roma-Mateo, Perez-Machado, Peiro-Chova, & Pallardó, Reference García-Giménez, Roma-Mateo, Perez-Machado, Peiro-Chova and Pallardó2017).
Although evidence examining this pathway in the context of epigenetics is scarce, some EWAS have shown interesting results: one study examined prospectively the association of EWAS methylation changes with the transition to psychosis (Kebir et al., Reference Kebir, Chaumette, Rivollier, Miozzo, Lemieux Perreault, Barhdadi and Krebs2017), and found an enrichment of pathways related to oxidative stress regulation in those transitioning. Furthermore, an EWAS study in Borderline PD found differences in methylation in GCT6, which is important in glutathione metabolism (Arranz et al., Reference Arranz, Gallego-Fabrega, Martín-Blanco, Soler, Elices, Dominguez-Clavé and Pascual2021).
The epigenetic signature of childhood adversity and cannabis use
The characteristic of the studies described in this section can be found in Online Supplementary Table S1 (SM), and in Table 2.
Hypothalamus pituitary-adrenal axis pathway
While multiple studies have explored the link between epigenetic modification involved in the HPA-axis, and psychiatric disorders, recent evidence is beginning to indicate that some of these epigenetic modifications might follow exposure to CA. The latter is a robustly replicated risk factors for many psychiatric disorders (Online Supplementary Table S1 (SM) summarises the main findings on studies examining the link between DNAm and genes involved in the HPA-axis, and key findings are summarised in Fig. 2). As a whole, as illustrated in Fig. 2, at the level of NR3C1 there is consistent evidence on a positive correlation between CA and DNAm in Borderline PD and MDD and some clinical outcomes (Farrell et al., Reference Farrell, Doolin, O'Leary, Jairaj, Roddy, Tozzi and Nemoda2018; Martin-Blanco et al., Reference Martin-Blanco, Ferrer, Soler, Salazar, Vega and Andion2014; Perroud et al., Reference Perroud, Paoloni-Giacobino, Prada, Olie, Salzmann and Nicastro2011; Radtke et al., Reference Radtke, Schauer, Gunter, Ruf-Leuschner, Sill, Meyer and Elbert2015), and a negative correlation with anxiety and PTSD (Labonte, Azoulay, Yerko, Turecki, & Brunet, Reference Labonte, Azoulay, Yerko, Turecki and Brunet2014; Schechter et al., Reference Schechter, Moser, Paoloni-Giacobino, Stenz, Gex-Fabry, Aue and Manini2015; Wang et al., Reference Wang, Feng, Ji, Mu, Ma, Fan and Zhu2017; Yehuda et al., Reference Yehuda, Flory, Bierer, Henn-Haase, Lehrner, Desarnaud and Meaney2015). Lower DNAm in FKBP5 is associated with CA in psychosis and PTSD (Klengel et al., Reference Klengel, Mehta, Anacker, Rex-Haffner, Pruessner and Pariante2013; Misiak et al., Reference Misiak, Karpiński, Szmida, Grąźlewski, Jabłoński, Cyranka and Frydecka2020); while in depression, 3 studies found no such link (Bustamante et al., Reference Bustamante, Aiello, Guffanti, Galea, Wildman and Uddin2018; Farrell et al., Reference Farrell, Doolin, O'Leary, Jairaj, Roddy, Tozzi and Nemoda2018; Klinger-König et al., Reference Klinger-König, Hertel, Van der Auwera, Frenzel, Pfeiffer, Waldenberger and Homuth2019), as opposed to another study (Tozzi et al., Reference Tozzi, Farrell, Booij, Doolin, Nemoda, Szyf and Frodl2018). As for NR3C1, findings on FKBP5 DNAm are different across disorder, suggesting a divergent transdiagnostic mechanism involving in HPA related genes (see Fig. 2). The SKA2 interacts with adversity scores in predicting lifetime suicide attempt (Kaminsky et al., Reference Kaminsky, Wilcox, Eaton, Van Eck, Kilaru, Jovanovic and Ressler2015), and mediated the association between reduced cortical thickness and PTSD (Sadeh et al., Reference Sadeh, Spielberg, Logue, Wolf, Smith, Lusk and McGlinchey2016a) and suicide phenotypes (Sadeh et al., Reference Sadeh, Wolf, Logue, Hayes, Stone, Griffin and Miller2016b).
Serotoninergic, dopaminergic and glutamatergic/GABAergic pathways
Childhood adversity
With regards to the serotoninergic pathway, while hypomethylation in SLC6A is associated with resilience to PTSD (Koenen et al., Reference Koenen, Uddin, Chang, Aiello, Wildman, Goldmann and Galea2011), hypermethylation of SLC6A has been linked to exposure to CA and associated with the worst clinical presentation in MDD (Kang et al., Reference Kang, Kim, Stewart, Kim, Bae, Kim and Yoon2013). Moreover, hypermethylation in the 5-HT3A-R gene appeared to mediate the link between exposure to adversity and higher severity of disease parameters in a mixed sample of BD and Borderline PD (Perroud et al., Reference Perroud, Zewdie, Stenz, Adouan, Bavamian and Prada2016).
Moreover, hypomethylation of MAOA, a gene important for the degradation of serotonin and DA (Xu, Jiang, Gu, Wang, & Yuan, Reference Xu, Jiang, Gu, Wang and Yuan2020), appears to partially mediate the known association between CA and depressive symptoms, alongside with other stress-related genes such as BDNF and NR3C1 and SLC64 (Peng et al., Reference Peng, Zhu, Strachan, Fowler, Bacus, Roy-Byrne and Zhao2018). Moreover, MAOA DNAm was negatively correlated to life events in a sample of Panic Disorder (Domschke et al., Reference Domschke, Tidow, Kuithan, Schwarte, Klauke, Ambrée and Kersting2012).
In relation to DA, one study in patients with bulimia spectrum disorders found no differences in DRD2 DNAm when comparing those with exposure and non-exposure to trauma (Groleau et al., Reference Groleau, Joober, Israel, Zeramdini, DeGuzman and Steiger2014).
At the level of the Glutamatergic pathway, one study found that exposure to CA was associated with decreased DNAm in GAD in a sample of Panic Disorder (Domschke et al., Reference Domschke, Tidow, Schrempf, Schwarte, Klauke, Reif and Deckert2013). Lastly, a candidate gene study (Engdahl, Alavian-Ghavanini, Forsell, Lavebratt, & Rüegg, Reference Engdahl, Alavian-Ghavanini, Forsell, Lavebratt and Rüegg2021) and an EWAS (Weder et al., Reference Weder, Zhang, Jensen, Yang, Simen, Jackowski and Perepletchikova2014) linked CA to increased methylation levels in GRIN2B/GRIND1 genes, suggesting evidence that this change may lead to the onset of depressive symptoms.
As a whole, DNAm changes in some of the genes that have been previously linked to major psychiatric conditions (Section ‘Evidence of epigenetic processes in major transdiagnostic pathways’, Table 2), such as SLC6A, 5HT3A-R, A (MAOA), BDNF, GAD and the GRIND family, (related to the serotoninergic, and glutamatergic pathways respectively) are also associated to CA. This suggests that some of the DNAm changes attributed to these disorders may be partially related to the consequence of CA exposure, as illustrated in Fig. 1.
Cannabis use
CU and in particular heavy use (Marconi et al., Reference Marconi, Di Forti, Lewis, Murray and Vassos2016) has been consistently associated with increased risk for PD, but to a lesser degree for other psychiatric conditions (Sideli, Trotta, Spinazzola, La Cascia, & Di Forti, Reference Sideli, Trotta, Spinazzola, La Cascia and Di Forti2020b). In recent years, candidate genes studies from peripheral blood have reported changes in DNAm associated with heavy CU in genes involved in dopamine transmission, such as DRD2 (Gerra et al., Reference Gerra, Jayanthi, Manfredini, Walther, Schroeder, Phillips and Donnini2018), DAT1 (Grzywacz et al., Reference Grzywacz, Barczak, Chmielowiec, Chmielowiec, Suchanecka, Trybek and Rubis2020) and COMT (Van der Knaap et al., Reference Van der Knaap, Schaefer, Franken, Verhulst, van Oort and Riese2014) and in the CB1 and CB2 receptors genes part of the endocannabinoid system (Rotter et al., Reference Rotter, Bayerlein, Hansbauer, Weiland, Sperling, Kornhuber and Biermann2012; Tao et al., Reference Tao, Li, Jaffe, Shin, Deep-Soboslay, Yamin and Kleinman2020). The latter playing an important role in brain development and synaptic transmission.
A recent study investigated the effect of heavy CU with and without tobacco on EWAS (Osborne et al., Reference Osborne, Pearson, Noble, Gemmell, Horwood, Boden and Kennedy2020). The analyses in the sample with both cannabis and tobacco use identified differentially methylated sites in 2 genes, AHRR and F2RL, previously reported to be affected by tobacco exposure. Within the sample of cannabis users without tobacco, while none of the differentially methylated loci reached EWAS significance, an exploratory analysis showed enrichment for genes involved in the signalling pathway, including glutamatergic transmission, brain function and mood disorders. Moreover, these exploratory analyses show two differentially methylated sites significantly associated with both only CU and cannabis with tobacco, which are within the MARC2 gene. The latter previously linked to adverse effects to antipsychotics in schizophrenia (Åberg et al., Reference Åberg, Adkins, Bukszár, Webb, Caroff, Miller and Vladimirov2010) and within the CUX1 gene which is involved in neuronal development (Platzer et al., Reference Platzer, Cogné, Hague, Marcelis, Mitter, Oberndorff and van der Smagt2018).
Furthermore, recent whole blood and cell-specific Methylome-wide association studies (MWAS) from a sample of adolescents with CU disorder pointed at many methylation sites relevant to brain function and to neurodevelopment (Clark et al., Reference Clark, Chan, Zhao, Xie, Copeland, Aberg and van den Oord2021). These included CpGs located in the CLMN gene and the SENP7 gene, expressed in the brain and playing a role in brain developmental and synaptic function and organisation (Juarez-Vicente, Luna-Pelaez, & Garcia-Dominguez, Reference Juarez-Vicente, Luna-Pelaez and Garcia-Dominguez2016; Marzinke & Clagett-Dame, Reference Marzinke and Clagett-Dame2012). Interestingly, the pathway analyses based on the cell type-specific significant DNAm changes associated with CU implicated pathways such as the Slit-Robo signalling (granulocytes) under the regulatory control of the endocannabinoid system during brain cortical development (Alpár et al., Reference Alpár, Tortoriello, Calvigioni, Niphakis, Milenkovic, Bakker and Fuzik2014), the ErbB signalling pathway (T-cell) and pathways involved in DNA repair (B-cell) (Clark et al., Reference Clark, Chan, Zhao, Xie, Copeland, Aberg and van den Oord2021).
Inflammation, oxidative stress, synaptic plasticity and neurogenesis
Childhood adversity
A number of EWAS studies conducted in clinical samples have reported an association between exposure to CA and DNAm changes across genes involved in inflammation. For instance, a study (Prados et al., Reference Prados, Stenz, Courtet, Prada, Nicastro and Adouan2015) found a positive correlation between the IL17RA DNAm and CA in a Borderline PD and MDD sample. Other evidence suggests a negative correlation between DNAm in genes enriched for immune pathways (such as TLR1 and TLR3) and CA in PTSD subjects (Uddin et al., Reference Uddin, Aiello, Wildman, Koenen, Pawelec, de Los Santos and Galea2010); while the TNFRSF13C gene was differently methylated between Borderline PD participants with and without CA (Arranz et al., Reference Arranz, Gallego-Fabrega, Martín-Blanco, Soler, Elices, Dominguez-Clavé and Pascual2021) (See Online Supplementary Table S1 (SM) – EWAS section).
Candidate genes studies have linked CA with DNAm changes in BDNF (Moser et al., Reference Moser, Paoloni-Giacobino, Stenz, Adouan, Manini, Suardi and Rusconi-Serpa2015; Thaler et al., Reference Thaler, Gauvin, Joober, Groleau, Guzman and Ambalavanan2014; Weder et al., Reference Weder, Zhang, Jensen, Yang, Simen, Jackowski and Perepletchikova2014), consistently with EWAS data reporting DNAm changes affecting genes involved in neurogenesis (Prados et al., Reference Prados, Stenz, Courtet, Prada, Nicastro and Adouan2015; Uddin et al., Reference Uddin, Aiello, Wildman, Koenen, Pawelec, de Los Santos and Galea2010). For instance, three EWAS studies in BD (Comes et al., Reference Comes, Andlauer, Adorjan, Budde, Gade, Degenhardt and Kondofersky2019), Borderline PD (Arranz et al., Reference Arranz, Gallego-Fabrega, Martín-Blanco, Soler, Elices, Dominguez-Clavé and Pascual2021) and MDD (Lutz et al., Reference Lutz, Tanti, Gasecka, Barnett-Burns, Kim, Zhou and Almeida2017) have consistently shown changes in DNAm in genes from the POU family that are associated with CA (POU6F2, POU5F1 and POU3F1 respectively), which are genes involved in myelinisation and neurogenesis (Online Supplementary Table S1 (SM) – EWAS section).
A recent EWAS study found differences in DNAm of the GGT6 gene that were associated with exposure to CA in a sample of Borderline PD patients; GGT6 is key for glutathione homoeostasis (Arranz et al., Reference Arranz, Gallego-Fabrega, Martín-Blanco, Soler, Elices, Dominguez-Clavé and Pascual2021), it is also the main antioxidant and redox regulator that has previously been associated with SCZ aetiopathogenesis (Steullet et al., Reference Steullet, Cabungcal, Monin, Dwir, O'Donnell, Cuenod and Do2016). Further evidence is summarised in Online Supplementary Table S2 (SM).
As a whole, candidate gene and EWAS studies suggest a link between CA and genes involved in the inflammatory and neurogenesis pathways, with some preliminary evidence suggesting a link between CA and DNAm and oxidative stress genes (Fig. 1).
Cannabis use
The largest to date case–control study to examine the effect of lifetime CU on DNAm reported an epigenome-wide-significantly differentially methylated CpG site within the CEMIP gene (Markunas et al., Reference Markunas, Hancock, Xu, Quach, Fang, Sandler and Taylor2020). The CEMIP gene, involved in hyaluronic catabolism, which has been shown to have an important role in inflammation, immune processes as well as associated with BD and SCZ previously (Petrey & de la Motte, Reference Petrey and de la Motte2014).
Other environmental exposures that can act as confounders in psychiatric epigenetic
Tobacco smoking
A number of publications have identified robust associations between tobacco smoking and DNAm (Elliott et al., Reference Elliott, Tillin, McArdle, Ho, Duggirala, Frayling and Relton2014; Shenker et al., Reference Shenker, Polidoro, van Veldhoven, Sacerdote, Ricceri, Birrell and Flanagan2013; Tsaprouni et al., Reference Tsaprouni, Yang, Bell, Dick, Kanoni, Nisbet and Deloukas2014; Zeilinger et al., Reference Zeilinger, Kuhnel, Klopp, Baurecht, Kleinschmidt, Gieger and Illig2013), with a number of genes (e.g. AHRR, F2RL3, GFI1 and MYO1G) replicated across studies.
The confounding effect of smoking is clearly evidenced in an EWAS study on peripheral blood of SCZ patients (Hannon et al., Reference Hannon, Dempster, Viana, Burrage, Smith, Macdonald and Mill2016). A similar study examining the impact of CA on the epigenome in a general population found that tobacco consumption was an important confounding when examining the signature of CA (Marzi et al., Reference Marzi, Sugden, Arseneault, Belsky, Burrage, Corcoran and Caspi2018). Whether some of these epigenetic changes associated with tobacco exposure could also mediate the already reported link between tobacco use and increased risk of psychosis, it is an important question yet to be determined (Gurillo et al., Reference Gurillo, Jauhar, Murray and MacCabe2015), and tobacco smoking should be accounted for in the future epigenetic studies in psychiatry.
Alcohol use and abuse
There is some initial evidence to suggest that alcohol use is associated with DNAm changes (Liu et al., Reference Liu, Marioni, Hedman, Pfeiffer, Tsai, Reynolds and Tanaka2016; Wang, Xu, Zhao, Gelernter, & Zhang, Reference Wang, Xu, Zhao, Gelernter and Zhang2016; Weng, Wu, Lee, Hsu, & Cheng, Reference Weng, Wu, Lee, Hsu and Cheng2015). Enrichment analyses examined DNAm in alcohol users have revealed enrichment in pathways related to neural degeneration (Weng et al., Reference Weng, Wu, Lee, Hsu and Cheng2015), and in genes important for neurogenesis (NPDC1), inflammation (HERC5) and in GABA receptors (a receptor delta and B receptor subunit 1); all of which are pathways previously associated with different mental disorders, as shown in Fig. 1. However, studies rarely account for such covariates, which is currently a limitation of current literature.
Psychiatric medication
The extent of the data reporting the DNAm changes associated with psychiatric medication would require a separate review. Indeed, there is consistent evidence that pharmacological agents can trigger DNAm in similar or opposite directions than those attributed to the disease. For example, Lithium, Carbamazepine and Quetiapine, often prescribed for the treatment of BD, are associated with decrease methylation of SLC6A4 (Asai et al., Reference Asai, Bundo, Sugawara, Sunaga, Ueda, Tanaka and Iwamoto2013; Sugawara et al., Reference Sugawara, Bundo, Asai, Sunaga, Ueda, Ishigooka and Iwamoto2015), in contrast with the hypermethylation reported in BD in that gene (Table 2). Similarly, studies who have investigated the effect of antipsychotic medication, have shown, on the one hand, that Haloperidol affects DNAm in leucocytes of SCZ patients (Melas et al., Reference Melas, Rogdaki, Ösby, Schalling, Lavebratt and Ekström2012), while on the other hand, a recent EWAS study showed that Clozapine exposure leads to DNAm differences in patients with treatment-resistant SCZ (Hannon et al., Reference Hannon, Dempster, Mansell, Burrage, Bass, Bohlken and Mill2021) as compared to controls. Thus, it is key to consider the possibility that some of the changes in DNA pathways may be led by agents rather than the disease itself, highlighting the need to account for medication in future studies and to consider epigenetics as a potential mediating mechanism of action of the beneficial effects of medication in the brain.
Summary and outstanding questions
As illustrated in Fig. 1, many of the epigenetic dysregulations we report are transdiagnostic, such as those affecting the serotoninergic, inflammatory and neurogenesis pathways, while others such as the Glutamatergic/GABAergic pathway are shared between a couple of disorders (e.g. SCZ and ASD), or disorder specific such as the dopaminergic pathway in PDs. These are pathways that have been classically implicated in the aetiopathogenesis of psychiatric phenotypes; additional emerging pathways such as oxidative stress remain to be further explored.
Moreover, CA, is transdiagnostically associated to psychiatry morbidity, and seems to play a role in the DNAm dysregulation of many of these pathways. Furthermore, the preliminary DNAm changes so far reported associated with CU affect pathways previously link to psychosis, suggesting potential mediating venues to be tested in clinical populations (Fig. 1).
In addition, CA is associated with DNAm changes both in the general population (Kandaswamy et al., Reference Kandaswamy, Hannon, Arseneault, Mansell, Sugden, Williams and Wong2020) as well as in clinical samples with a psychiatric diagnosis (Online Supplementary Table S1 (SM)). This might suggest that the DNAm changes associated to CA exposure predate disease onset and could represent a marker of acquired psychiatric liability. However, evidence formally testing mediating pathways EWAS level between CA and the main clinical conditions is non-existent in humans. Candidate gene studies tend to find the inconsistent direction of the association between CA across disorders, or findings are inconsistent within disorders as shown in Fig. 2 and Table 2 and Online Supplementary Table S1 (SM). One explanation could be that there are other causative partners that are being missed in the equation, that may explain such inconsistency, such as the role of genotype, gene expression or a more thorough assessment of specific adversities in the context of protective factors and its link with more carefully selected clinical phenotypes.
The existing findings from epigenetics research need to be appraised in the context of well-known technical limitations epigenetics, such as the blood-brain inconsistencies, tissue-type specificity (Bakulski, Halladay, Hu, Mill, & Fallin, Reference Bakulski, Halladay, Hu, Mill and Fallin2016; Nikolova & Hariri, Reference Nikolova and Hariri2015) and the candidate gene v. EWAS issue (see Palma-Gudiel, Córdova-Palomera, Leza, & Fañanás, Reference Palma-Gudiel, Córdova-Palomera, Leza and Fañanás2015). Moreover, evidence suggests that variation in DNAm depends not only on the environment, but also on genetic factors (Bell & Spector, Reference Bell and Spector2012). Although some studies presented in this review have found evidence that some genotypic variation in some risk alleles can influence DNAm (Klengel et al., Reference Klengel, Mehta, Anacker, Rex-Haffner, Pruessner and Pariante2013; Melas et al., Reference Melas, Wei, Wong, Sjöholm, Åberg, Mill and Lavebratt2013; Perroud et al., Reference Perroud, Zewdie, Stenz, Adouan, Bavamian and Prada2016), EWAS addressing the joint effect of genotype and environment are still in its infancy (Min et al., Reference Min, Hemani, Hannon, Dekkers, Castillo-Fernandez, Luijk and Suderman2021). Addressing this issue will prove methodologically challenging, but methods quantifying the genetic influences on DNAm, such as the methylation quantitative trait loci (mQTL) should be used in relation to the presence of environmental insults. Moreover, studies included in this work are often small (Online Supplementary Table S1 (SM)) and thus underpowered, except some exceptions (Hannon et al., Reference Hannon, Dempster, Mansell, Burrage, Bass, Bohlken and Mill2021), which presents the need to create collaborative efforts allowing meta-analysis of comparable epigenetic data. Furthermore, evidence of the environmental exposure impact through epigenetic modification in psychiatric diseases or phenotypes is still limited, with studies focusing mainly on exposure to CA and only preliminary results of the effect of cannabis. Given the replicated but differential impact of multiple environmental risk factors in major psychiatric disorders (Rodriguez et al., Reference Rodriguez, Alameda, Trotta, Spinazzola, Marino, Matheson and Vassos2021), future studies exploring epigenetic variation as a mediator between genetic vulnerability and various environmental factors (not only CA) should be addressed, using novel methods specifically developed for mediation using EWAS data (Liu et al., Reference Liu, Shen, Barfield, Schwartz, Baccarelli and Lin2021). Another important factor is the phenotypic characterisation for environmental exposure. For instance, most of the studies in this work have used broad measures of adversity, using a composite cumulative score, rather than differentiating between neglect of abuse. The same can be said for the measures of CU which little reflects the level of exposure none to affect psychiatric liability. Moreover, the outcomes are often considered as SCZ, or MDD or even major psychoses (combining SCZ & BD), which are extremely heterogeneous entities, involving microphenotypes (Maj et al., Reference Maj, van Os, De Hert, Gaebel, Galderisi, Green and Malaspina2021), and which accordingly may have very different biological underpinnings. Evidence is showing that there are some levels of specificity between adversity subtypes and symptoms domains, for example, abuse is more related to positive symptoms while neglect is not (Alameda et al., Reference Alameda, Christy, Rodriguez, Salazar de Pablo, Thrush, Shen and Murray2021) and that CU is associated with paranoia (Freeman et al., Reference Freeman, Garety, Bebbington, Smith, Rollinson, Fowler and Dunn2005). Thus, using a composite measure of CA and broadly defined conditions when trying to understand specific mediating epigenetic pathways may consider such specific links between environment and psychopathology first. Accordingly, this work suggests that some biological pathways are operating transdiagnostically, and therefore a phenotypic characterisation based on clinical dimensions may be more biologically informative than diagnostic categorisations. Furthermore, the timing of environmental exposure should be addressed, given evidence that a disruption in epigenetic programming occurs across different time windows throughout the life span (Massicotte, Whitelaw, & Angers, Reference Massicotte, Whitelaw and Angers2011). In this line, the lack of information on the timing of trauma and of CU initiation could explain some of the inconsistencies mentioned in our review (Fig. 2). For example, we reviewed studies showing increased methylation of the serotonin transporter in depressed individuals exposed to trauma (usually when adversity occurs before adulthood), which contrasts with the lower methylation in the same gene in PTSD, when exposure tends to be later in life.
Conclusions
Future Research should include the influence of gender and how it can modulate the links between DNAm and mental disorders, or how it can affect the influence of CA on DNAm. More effort should go towards designing studies that integrate genetic data with the often-neglected effect of environmental exposures (e.g. recreational drugs and psychotropic medication). Specifically, collaborative efforts between geneticists, epigeneticists and epidemiologists will lead to increased understanding of how the DNAm changes mapping to specific pathways, might mediate the biological link between environmental exposures and increased liability to specific or transdiagnostic psychiatric morbidity.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0033291721005559.
Acknowledgements
We thank our funding bodies. Dr Luis Alameda was supported at the beginning of the preparation of the current work by the Swiss national Foundation (P2LAP3 171804). Dr Chloe Wong is supported by a joint grant from the Economic and Social Research Council (ESRC) and Biotechnology and Biological Sciences Research Council [ES/N000277/1]. Dr Marta Di Forti work is supported by the Medical Research Council SRF MR/T007818/1.







