INTRODUCTION
Giardia duodenalis (syn. Giardia intestinalis and Giardia lamblia), is a protozoan enteric parasite that causes acute, watery diarrhoea or giardiasis in 280 million people annually and is a common cause of waterborne outbreaks (Lane and Lloyd, Reference Lane and Lloyd2002; Baldursson and Karanis, Reference Baldursson and Karanis2011; Painter et al. Reference Painter, Gargano, Collier and Yoder2015; Einarsson et al. Reference Einarsson, Ma'ayeh and Svärd2016). Most infections are self-limiting but chronic infections can lead to weight loss and malabsorption (Ryan and Cacciò, Reference Ryan and Cacciò2013) and infections are associated with stunting (low height for age), wasting (low weight for height) and cognitive impairment in children in developing countries (Berkman et al. Reference Berkman, Lescano, Gilman, Lopez and Black2002; Feng and Xiao, Reference Feng and Xiao2011). Furthermore, acute giardiasis may disable patients for extended periods and can elicit protracted post-infectious syndromes, including irritable bowel syndrome and chronic fatigue (Hanevik et al. Reference Hanevik, Wensaas, Rortveit, Eide, Mørch and Langeland2014). In Australia, the overall prevalence is ~2 to 7%, with a steady increase of reports each year (Asher et al. Reference Asher, Hose and Power2016). Giardia is responsible for ~614, 740 sporadic cases of acute gastroenteritis per year in Australia and a disease burden of 4·3 disability-adjusted life years (Gibney et al. Reference Gibney, O'Toole, Sinclair and Leder2014).
Giardia duodenalis consists of eight genetic assemblages (A–H) with different host specificities; assemblage A in humans, livestock and other mammals; B in humans, primates and some other mammals, C and D in dogs and other canids; E mainly in hoofed animals including cattle, sheep and goats, and more recently in rabbits, non-human primates and humans; F in cats and humans, and more recently in cattle; G in rats and mice; and H in marine mammals (Andrews et al. Reference Andrews, Adams, Boreham, Mayrhofer and Meloni1989; Mayrhofer et al. Reference Mayrhofer, Andrews, Ey and Chilton1995; Ey et al. Reference Ey, Mansouri, Kulda, Nohýnková, Monis, Andrews and Mayrhofer1997; Monis et al. Reference Monis, Andrews, Mayrhofer, Kulda, Isaac-Renton and Ey1998; Ryan and Cacciò, Reference Ryan and Cacciò2013; Cardona et al. Reference Cardona, de Lucio, Bailo, Cano, de Fuentes and Carmena2015; Qi et al. Reference Qi, Xi, Li, Wang, Ning and Zhang2015). Of these, assemblages A and B are the predominant assemblages in humans and exhibit a broad host range including cattle, sheep, pigs, horses, non-human primates, dogs, cats and fish (Yang et al. Reference Yang, Reid, Lymbery and Ryan2010a; Feng and Xiao, Reference Feng and Xiao2011; Ghoneim et al. Reference Ghoneim, Abdel-Moein and Saeed2012; Ryan and Cacciò, Reference Ryan and Cacciò2013; Durigan et al. Reference Durigan, Abreu, Zucchi, Franco and de Souza2014). The remaining assemblages are considered to be largely host-specific. However recent studies have reported the occurrence of animal assemblages in humans including assemblage F in Ethiopia (Gelanew et al. Reference Gelanew, Lalle, Hailu, Pozio and Cacciò2007), assemblage E in Egypt and Brazil (Foronda et al. Reference Foronda, Bargues, Abreu-Acosta, Periago, Valero, Valladares and Mas-Coma2008; Helmy et al. Reference Helmy, Klotz, Wilking, Krücken, Nöckler, Samson-Himmelstjerna, Zessin and Aebischer2014; Abdel-Moein and Saeed, Reference Abdel-Moein and Saeed2016; Fantinatti et al. Reference Fantinatti, Bello, Fernandes and Da-Cruz2016; Scalia et al. Reference Scalia, Fava, Soares, Limongi, da Cunha, Pena, Kalapothakis and Cury2016), assemblage C in China and Slovakia (Liu et al. Reference Liu, Shen, Yin, Yuan, Jiang, Xu, Pan, Hu and Cao2014; Štrkolcová et al. Reference Štrkolcová, Maďar, Hinney, Goldová, Mojžišová and Halánová2015) and assemblage D in German travellers (Broglia et al. Reference Broglia, Weitzel, Harms, Cacció and Nöckler2013). Very little is known about the prevalence and genetic diversity of Giardia assemblages infecting humans in Queensland, Australia. To date, only two studies have investigated the molecular characteristics of Giardia species in human patients from Queensland (Nolan et al. Reference Nolan, Jex, Upcroft, Upcroft and Gasser2011; Ebner et al. Reference Ebner, Koehler, Robertson, Bradbury, Jex, Haydon, Stevens, Norton, Joachim and Gasser2015), with the latter reporting a prevalence of 1·9% for Northern Queensland and Darwin, while a prevalence of up to 12% in Queensland patients has been reported using microscopy (Boreham and Phillips, Reference Boreham and Phillips1986). The aim of the present study therefore was to examine G. duodenalis assemblages in human patients suffering from sporadic cases of giardiasis in order to better understand the epidemiology and transmission dynamics of this ubiquitous parasite in Queensland.
MATERIALS AND METHODS
Samples and DNA extraction
A total of 88 faecal samples diagnosed as Giardia-positive by microscopy (collected from patients from rural and urban areas in South-East Queensland) were shipped to Murdoch University by Department of Health (Queensland) staff under Murdoch University Human Ethics permit 2014/159. All samples were stored at 4 °C until analysed. Genomic DNA was extracted from 250 mg of each faecal sample using a Power Soil DNA Kit (MO BIO, Carlsbad, California). A negative control (no faecal sample) was used in each extraction group.
Quantitative polymerase chain reaction (qPCR) determination of cysts per gram of feces
All samples were initially amplified by qPCR analysis of the glutamate dehydrogenase (gdh) locus as previously described (Yang et al. Reference Yang, Jacobson, Gardner, Carmichael, Campbell and Ryan2014). Copy numbers detected were converted to cyst numbers on the basis that the gdh gene in Giardia is a single copy gene (Yee and Denis, Reference Yee and Dennis1992) and the fact that there are 4 haploid nuclei per cyst. Therefore, every 4 copies of gdh detected by qPCR were equivalent to 1 cyst.
Molecular typing
All samples were typed at the triose phosphate isomerase (tpi) locus using assemblage A, B and E specific primers as previously described (Sulaiman et al. Reference Sulaiman, Fayer, Bern, Gilman, Trout, Schantz, Das, Lal and Xiao2003; Geurden et al. Reference Geurden, Geldhof, Levecke, Martens, Berkvens, Casaert, Vercruysse and Claerebout2008; Levecke et al. Reference Levecke, Geldhof, Claerebout, Dorny, Vercammen, Cacciò, Vercruysse and Geurden2009). A representative subset of samples were also amplified using a nested PCR and sequenced at the glutamate dehydrogenase (gdh) locus (n = 30) as described by Read et al. (Reference Read, Monis and Thompson2004) and at the tpi (n = 27) locus using the assemblage-specific primers described above. Briefly, the amplified DNA from secondary tpi and gdh PCR products were separated by gel electrophoresis and purified for sequencing using an in-house filter tip method (Yang et al. Reference Yang, Murphy, Song, Ng-Hublin, Estcourt, Hijjawi, Chalmers, Hadfield, Bath, Gordon and Ryan2013). Purified PCR products were sequenced independently using an ABI Prism™ Dye Terminator Cycle Sequencing kit (Applied Biosystems, Foster City, California) according to the manufacturer's instructions at 60 °C annealing temperature for PCR products at the gdh locus and 64, 62 and 67 °C annealing temperature for assemblages A, B and E tpi PCR products, respectively (Read et al. Reference Read, Monis and Thompson2004; Geurden et al. Reference Geurden, Geldhof, Levecke, Martens, Berkvens, Casaert, Vercruysse and Claerebout2008; Levecke et al. Reference Levecke, Geldhof, Claerebout, Dorny, Vercammen, Cacciò, Vercruysse and Geurden2009). Sanger sequencing chromatogram files were imported into Geneious Pro 8·1·6 (Kearse et al. Reference Kearse, Moir, Wilson, Stones-Havas, Cheung, Sturrock, Buxton, Cooper, Markowitz, Duran, Thierer, Ashton, Meintjes and Drummond2012), where they were analysed, edited and aligned with reference sequences from GenBank using MUSCLE (Edgar, Reference Edgar2004) and trimmed.
Phylogenetic analysis
Phylogenetic analysis for both loci was conducted using Distance and Maximum Likelihood (ML) analysis in MEGA6 (after selection of the best nucleotide substitution models) (Tamura et al. Reference Tamura, Stecher, Peterson, Filipski and Kumar2013). Distance estimation was conducted based on evolutionary distances calculated with the Tamura-Nei model and grouped using Neighbour-Joining (NJ). ML was conducted using the Tamura 3-parameter model. The confidence of groupings was assessed by bootstrapping, using 1000 replicates.
RESULTS
Giardia duodenalis assemblage prevalences
Assemblage A and B were detected in 50% (44/88) and 38·6% (34/88) of samples respectively. Mixed A and B assemblages were identified in 4·5% (4/88) and assemblage E in 6·8% (6/88) of samples (Table 1). Information on age at the time of collection was only available for a subset of samples (n = 25) and of these, ages ranged from 3 months to 79 years. Information for three of the samples typed as assemblage E was available (Qld86, Qld81 and Qld82) and their ages were 11, 19 and 67 years, respectively and were from urban (Qld86, Qld81) and rural locations. Information on contact with animals was not available.
Table 1. Giardia duodenalis assemblages and sub-assemblages (where available) and cysts per gram of feces detected in Giardia-positive human faecal samples from Queensland, Australia
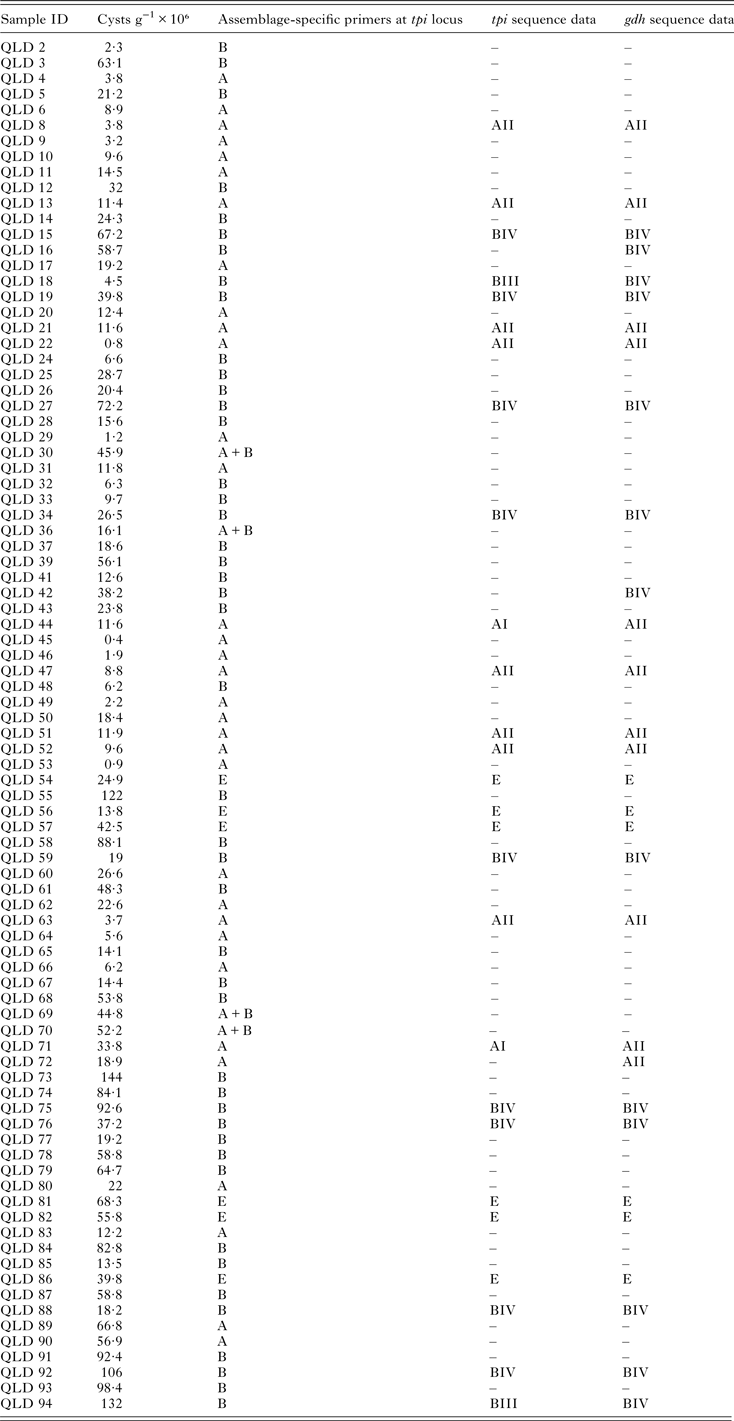
N/A, not available; “–”, not sequenced.
Molecular typing at the gdh and tpi loci
Phylogenetic analysis at the gdh and tpi loci using both Distance and ML analysis produced trees of similar topology (data not shown) and therefore only NJ trees are shown. Assignment to assemblage based on sequence analysis of the gdh and tpi loci was 100% concordant at the assemblage level between the two loci (i.e. all isolates typed as assemblage A by assemblage-specific PCR and sequence analysis of the tpi locus were also assemblage A at the gdh locus (Table 1). There were however differences at the sub-assemblage level. For example, at the gdh locus, all the assemblage A isolates that were typed (n = 11) were AII and exhibited 100% similarity to a Queensland human AII subtype from a Brisbane patient (AY178737) and an AII subtype from an Australian sheep (KY083429) (Fig. 1 – NJ tree and Table 1). At the tpi locus, two isolates (Qld 44 and Qld71) were typed as AI, while the remaining assemblage A isolates (n = 8) were typed as AII (Fig. 2 – NJ tree and Table 1). At the gdh locus, four of the assemblage B isolates (Qld16, Qld18, Qld42 and Qld94) grouped with an Australian reference BIV subtype (L40508), but exhibited 5 single nucleotide polymorphisms (SNP's) from the reference subtype across 666 bp of sequence. The remaining assemblage B isolates also grouped in the BIV clade but exhibited 7 SNP's with isolate VANC/94/UBC/125 (KP687771), across 699 bp of sequence. At the tpi locus, two isolates (Qld18 and Qld94) grouped with sub-assemblage BIII (100% similarity across 355 bp of sequence), while the remaining assemblage B isolates (n = 9) were 100% identical to a BIV sub-assemblage from a foal (KM926543) across 319 bp of sequence. At both the tpi and gdh locus, six isolates (Qld54, Qld56, Qld57, Qld81, Qld82, and Qld86) grouped in a clade with assemblage E. At the gdh locus, all six were 100% identical to a Western Australian (WA) cattle-derived assemblage E isolate (HQ398327), while at the tpi locus, all six were identical to a sheep-derived assemblage E isolate from Victoria (Vic) (GQ444454). Representative sequences have been submitted to GenBank under accession numbers KY655475- KY655485.
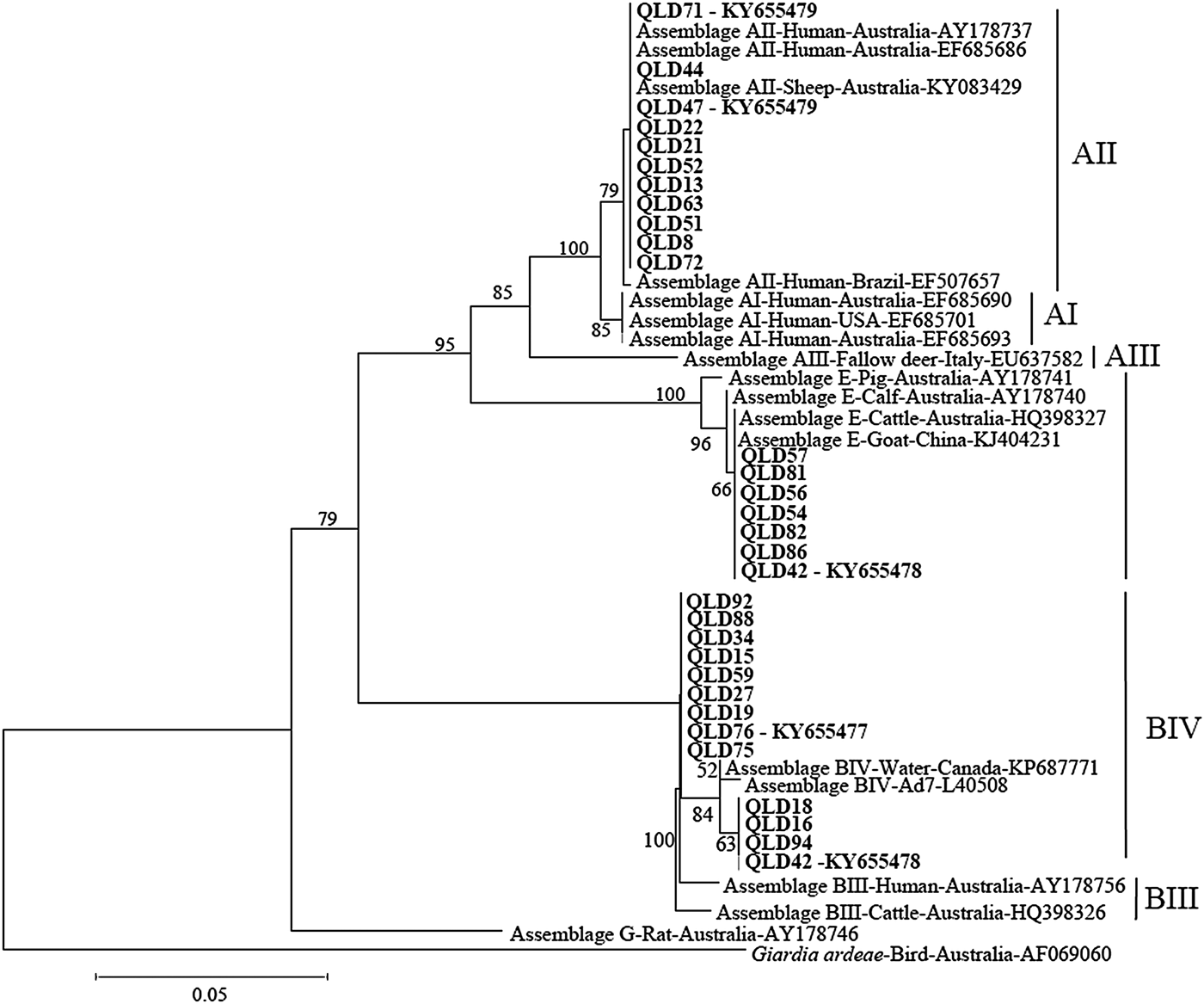
Fig. 1. Phylogenetic analysis of partial Giardia glutamate dehydrogenase (gdh) sequences amplified from human faecal samples inferred from Neighbour-Joining (NJ) analysis of Kimura's distances calculated from pair-wise comparisons. Percentage support (>50%) from 1000 pseudoreplicates from NJ analyses is indicated at the left of the supported node. Isolates from this study are indicated in bold font.
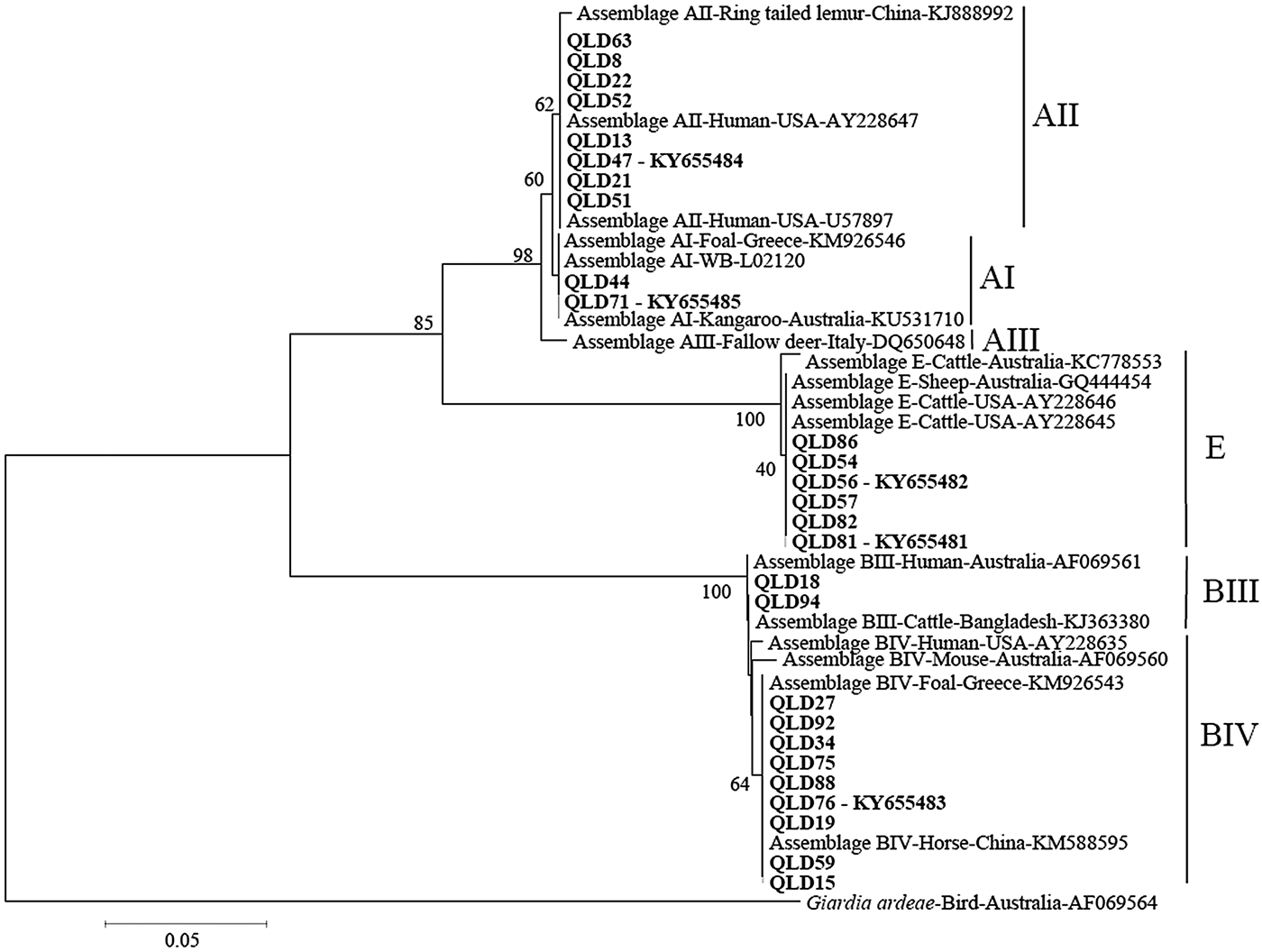
Fig. 2. Phylogenetic analysis of partial Giardia triose phosphate isomerase (tpi) sequences amplified from human faecal samples inferred from Neighbour-Joining (NJ) analysis of Kimura's distances calculated from pair-wise comparisons. Percentage support (>50%) from 1000 pseudoreplicates from NJ analyses is indicated at the left of the supported node. Isolates from this study are indicated in bold font.
Cysts per gram of feces
Cyst numbers per gram of feces (g−1) were determined using qPCR and ranged from 0·8 × 106 to 106 × 106 cysts g−1 (Table 1). Of the isolates that were typed as assemblage E, cyst numbers ranged from 13·8 to 68·3 × 106 cysts g−1.
DISCUSSION
In the present study, assemblage A and B were detected in 50% (44/88) and 38·6% (34/88) of samples, respectively. Mixed A and B assemblages were identified in 4·5% (4/88) and assemblage E in 6·8% (6/88) of samples. The predominance of assemblage A over B is in contrast with the findings from other studies of Giardia in humans in Australia. Previous studies have reported ~70–75% assemblage B and 25–30% assemblage A (n = 23–124) in Western Australia (Read et al. Reference Read, Walters, Robertson and Thompson2002; Yang et al. Reference Yang, Lee, Ng and Ryan2010b), 69% B (n = 9) and 31% A (n = 4) in patients from parts of the tropical North of Australia (Ebner et al. Reference Ebner, Koehler, Robertson, Bradbury, Jex, Haydon, Stevens, Norton, Joachim and Gasser2015) and 86·1% B and 12·7% A (n = 165) in New South Wales (Asher et al. Reference Asher, Hose and Power2016). Current evidence indicates that, on a global level, assemblage B is slightly more prevalent than assemblage A in both developed and developing countries (Feng and Xiao, Reference Feng and Xiao2011). The only two previous studies, which analysed a small number of Giardia isolates from Queensland, identified both assemblage A and B (Nolan et al. Reference Nolan, Jex, Upcroft, Upcroft and Gasser2011; Ebner et al. Reference Ebner, Koehler, Robertson, Bradbury, Jex, Haydon, Stevens, Norton, Joachim and Gasser2015).
Analysis of genetic variability within assemblages has shown that isolates of assemblage A can be divided into four sub-assemblages (AI, AII, AIII and AIV) by protein polymorphisms of 23 loci (Monis et al. Reference Monis, Mayrhofer, Andrews, Homan, Limper and Ey1996, Reference Monis, Andrews, Mayrhofer and Ey2003), with human isolates belonging to AI and AII and animal isolates belonged to AI, AIII and AIV (Monis et al. Reference Monis, Andrews, Mayrhofer and Ey2003; Cacciò et al. Reference Cacciò, Beck, Lalle, Marinculic and Pozio2008; Sprong et al. Reference Sprong, Cacciò and van der Giessen2009; Ryan and Cacciò, Reference Ryan and Cacciò2013). In the present study, although there was 100% concordance between loci in assignment to assemblage, there were some differences in assignment to sub-assemblage level. Of the subset of assemblage A isolates that were sequenced at the gdh locus (n = 30), all were AII, while at the tpi locus, two of the 10 assemblage A isolates were typed as AI. Sub-assemblages AI and AII are found in both humans and animals, with sub-assemblage AI preferentially found in livestock and pets, whereas sub-assemblage AII is predominantly found in humans (Sprong et al. Reference Sprong, Cacciò and van der Giessen2009). Previous studies in Australia have also reported that AII is the predominant sub-assemblage in humans (Yang et al. Reference Yang, Lee, Ng and Ryan2010b; Asher et al. Reference Asher, Hose and Power2016). It is also the dominant sub-assemblage in sheep across Australia (Yang et al. Reference Yang, Jacobson, Gardner, Carmichael, Campbell and Ryan2014) and therefore zoonotic transmission of AII is possible.
Within assemblage B, sub-assemblages BI, BII, BIII and BIV have been described by enzyme electrophoresis, with human isolates forming two clusters (BIII and BIV) and animal isolates (monkeys and a dog) belonging to sub-assemblages BI and BII (Monis et al. Reference Monis, Andrews, Mayrhofer and Ey2003). However, BIII and BIV sub-assemblages identified by allozyme electrophoresis are not always supported by DNA sequence analysis as subtyping analyses of field isolates produced inconsistent sub-assemblages among different loci (Feng and Xiao, Reference Feng and Xiao2011). In the present study, all the assemblage B isolates typed at the gdh locus (n = 13) are considered BIV variants, whereas at the tpi locus, two of the 11 assemblage B isolates were typed as BII, with the remainder BIV. Lack of concordance in the assignment to assemblages and sub-assemblages has been reported in many recent studies (Cacciò et al. Reference Cacciò, Beck, Lalle, Marinculic and Pozio2008; Ryan and Cacciò, Reference Ryan and Cacciò2013) and is explained by either mixed infections or recombination (Cacciò and Sprong, Reference Cacciò and Sprong2010). In the first case, two different assemblages/sub-assemblages are responsible for the infection of a single host and their detection depends on the relative proportion (with the majority population being favoured by PCR) and on the lack of bias during amplification. In the second case, genetic exchanges within or among assemblages can create recombinants that may contain specific sequences of assemblage A in a genome of assemblage B (or recombinants within sub-assemblages).
At both the gdh and tpi loci, six isolates were typed as assemblage E. At the gdh locus, all six were 100% identical to a WA cattle-derived assemblage E isolate (HQ398327) and at the tpi locus, were 100% identical to a Victorian sheep isolate (GQ444454). This is the first report of assemblage E in humans in Australia. A previous study in Egypt reported a high prevalence of assemblage E – 62·5% (25/40) in children living in agricultural areas in Egypt (Abdel-Moein and Saeed, Reference Abdel-Moein and Saeed2016). In that study, assemblage E was detected in 42·1% of Giardia-positive diarrheic and 81% of non-diarrheic children suggesting that assemblage E may cause clinical giardiasis (Abdel-Moein and Saeed, Reference Abdel-Moein and Saeed2016). The high prevalence of assemblage E was attributed to the fact that the children lived in rural villages with large cattle populations (Abdel-Moein and Saeed, Reference Abdel-Moein and Saeed2016). Other studies in Egypt have reported assemblage E in 15 and 11·1%, respectively, of Giardia positive human samples (Foronda et al. Reference Foronda, Bargues, Abreu-Acosta, Periago, Valero, Valladares and Mas-Coma2008; Helmy et al. Reference Helmy, Klotz, Wilking, Krücken, Nöckler, Samson-Himmelstjerna, Zessin and Aebischer2014). In Brazil, assemblage E was identified in 34% (15/44) of Giardia-positive samples amongst preschoolers (aged between 10 months and 4 years) in a community of Rio de Janeiro (Fantinatti et al. Reference Fantinatti, Bello, Fernandes and Da-Cruz2016). In that study, all samples were collected from children attending a day-care unit located in the slum with no sewerage network coverage, and stray animals including pigs and cattle moving throughout the location (Fantinatti et al. Reference Fantinatti, Bello, Fernandes and Da-Cruz2016).
Assemblages A and E are common amongst sheep and cattle in Australia with assemblage E the most dominant assemblage in hoofed animals (Nolan et al. Reference Nolan, Jex, Pangasa, Young, Campbell, Stevens and Gasser2010; Ng et al. Reference Ng, Yang, McCarthy, Gordon, Hijjawi and Ryan2011; Abeywardena et al. Reference Abeywardena, Jex, Firestone, McPhee, Driessen, Koehler, Haydon, von Samson-Himmelstjerna, Stevens and Gasser2013; Yang et al. Reference Yang, Jacobson, Gardner, Carmichael, Campbell and Ryan2014; Asher et al. Reference Asher, Hose and Power2016). In the present study, the individuals that were positive for Assemblage E were experiencing diarrhoea, came from both rural and urban areas and shed variable levels of cysts in their feces. All were identical to assemblage E from Australian cattle and sheep, suggesting possible zoonotic transmission. The data generated demonstrates that zoonotic transmission from cattle and sheep may be occurring and warrants further investigation.
ACKNOWLEDGEMENT
The authors wish to acknowledge the assistance of Frances Brigg and the Western Australia State Agriculture Biotechnology Centre for Sanger sequencing.
FINANCIAL SUPPORT
This study was financially supported by an Australian Research Council Linkage Grant number LP130100035.





