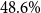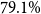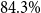A vast amount of clinical data are still stored in unstructured text. Automatic extraction of medical information from these data poses several challenges: high costs of clinical expertise, restricted computational resources, strict privacy regulations, and limited interpretability of model predictions. Recent domain adaptation and prompting methods using lightweight masked language models showed promising results with minimal training data and allow for application of well-established interpretability methods. We are first to present a systematic evaluation of advanced domain-adaptation and prompting methods in a lower-resource medical domain task, performing multi-class section classification on German doctor’s letters. We evaluate a variety of models, model sizes (further-pre)training and task settings, and conduct extensive class-wise evaluations supported by Shapley values to validate the quality of small-scale training data and to ensure interpretability of model predictions. We show that in few-shot learning scenarios, a lightweight, domain-adapted pretrained language model, prompted with just 20 shots per section class, outperforms a traditional classification model, by increasing accuracy from 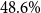 $48.6\%$ to
$48.6\%$ to 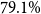 $79.1\%$. By using Shapley values for model selection and training data optimization, we could further increase accuracy up to
$79.1\%$. By using Shapley values for model selection and training data optimization, we could further increase accuracy up to 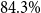 $84.3\%$. Our analyses reveal that pretraining of masked language models on general-language data is important to support successful domain-transfer to medical language, so that further-pretraining of general-language models on domain-specific documents can outperform models pretrained on domain-specific data only. Our evaluations show that applying prompting based on general-language pretrained masked language models combined with further-pretraining on medical-domain data achieves significant improvements in accuracy beyond traditional models with minimal training data. Further performance improvements and interpretability of results can be achieved, using interpretability methods such as Shapley values. Our findings highlight the feasibility of deploying powerful machine learning methods in clinical settings and can serve as a process-oriented guideline for lower-resource languages and domains such as clinical information extraction projects.
$84.3\%$. Our analyses reveal that pretraining of masked language models on general-language data is important to support successful domain-transfer to medical language, so that further-pretraining of general-language models on domain-specific documents can outperform models pretrained on domain-specific data only. Our evaluations show that applying prompting based on general-language pretrained masked language models combined with further-pretraining on medical-domain data achieves significant improvements in accuracy beyond traditional models with minimal training data. Further performance improvements and interpretability of results can be achieved, using interpretability methods such as Shapley values. Our findings highlight the feasibility of deploying powerful machine learning methods in clinical settings and can serve as a process-oriented guideline for lower-resource languages and domains such as clinical information extraction projects.